Andrew White | FutureHouse, Edison Scientific
Automating Scientific Discovery at Scale
Andrew White
FutureHouse, Edison Scientific
Altos Labs
April 2026
About Me
- 2015 - Assistant Professor at Rochester
- 2020 - Received Tenure
- 2023 - Cofounded FutureHouse, AIxBio non-profit
- 2025 - Cofounded Edison Scientific, AIxBio company
Science is changing independent of AI
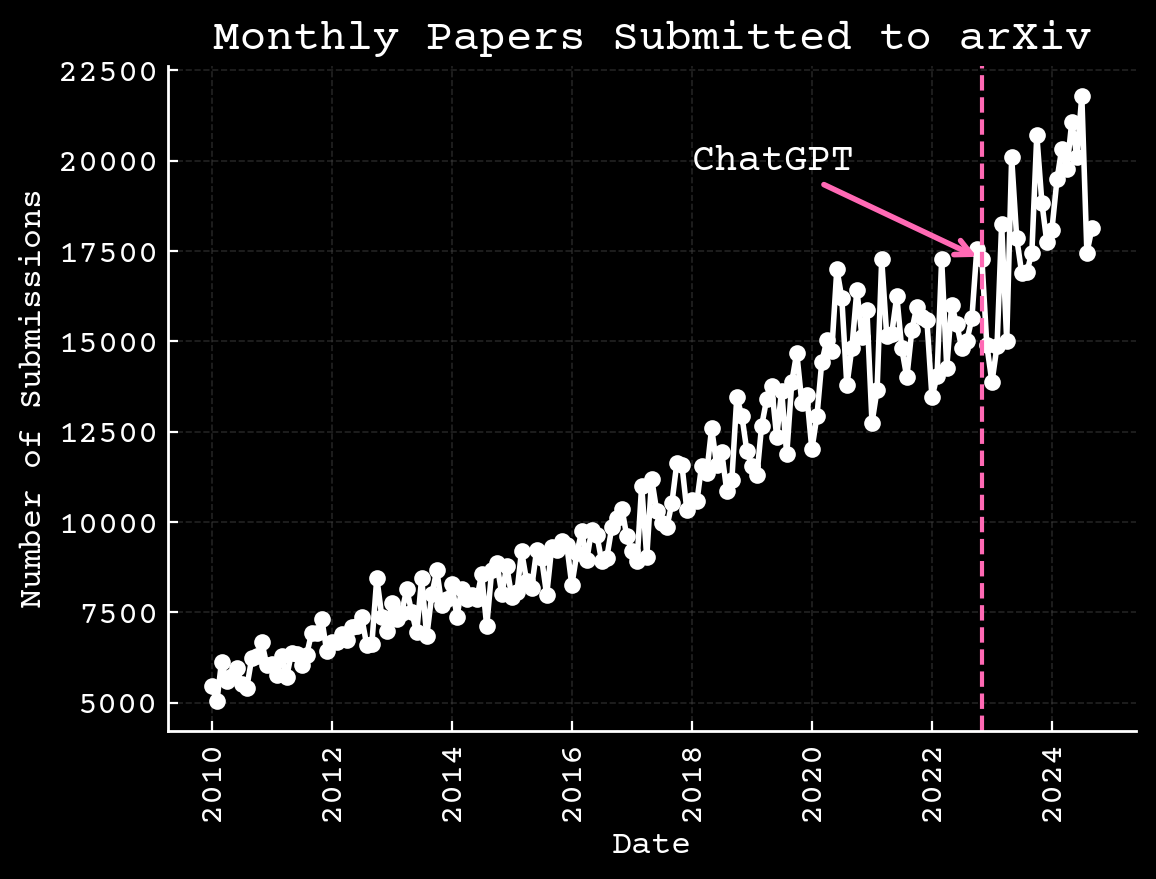
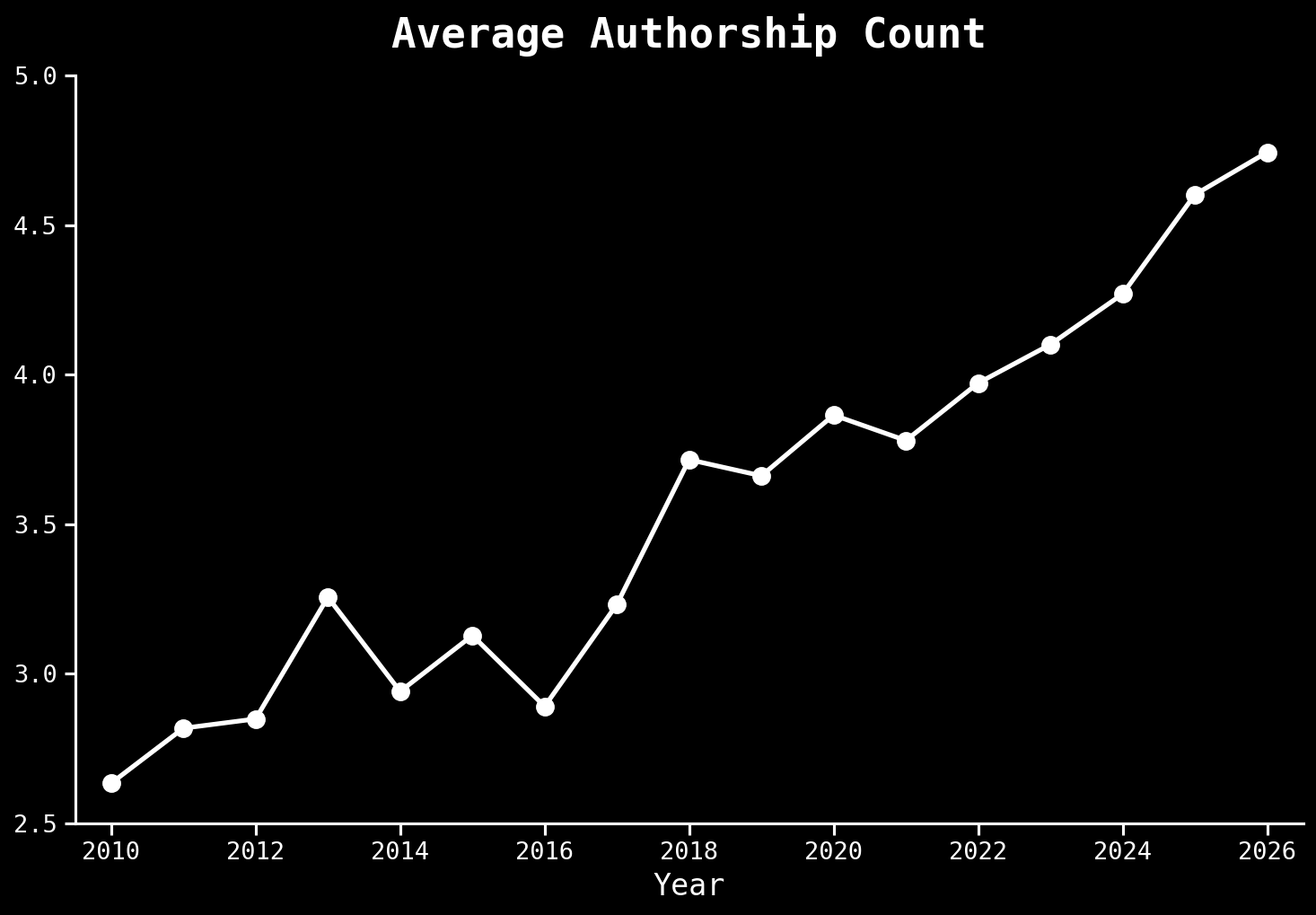
Arxiv.org,10.6084/m9.figshare.17064419.v3
Number of Researchers are Growing
International R&D spending
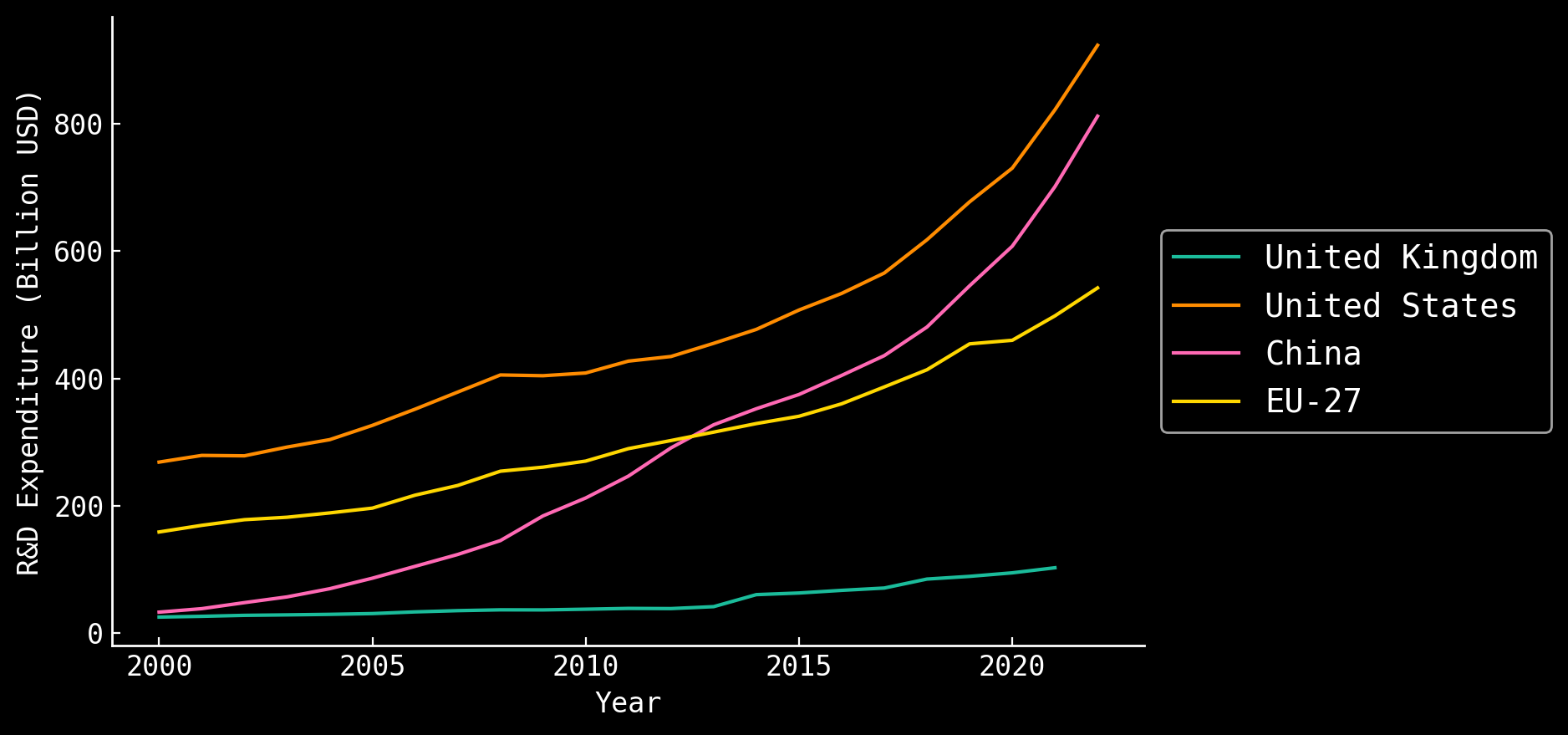
PhD Researchers
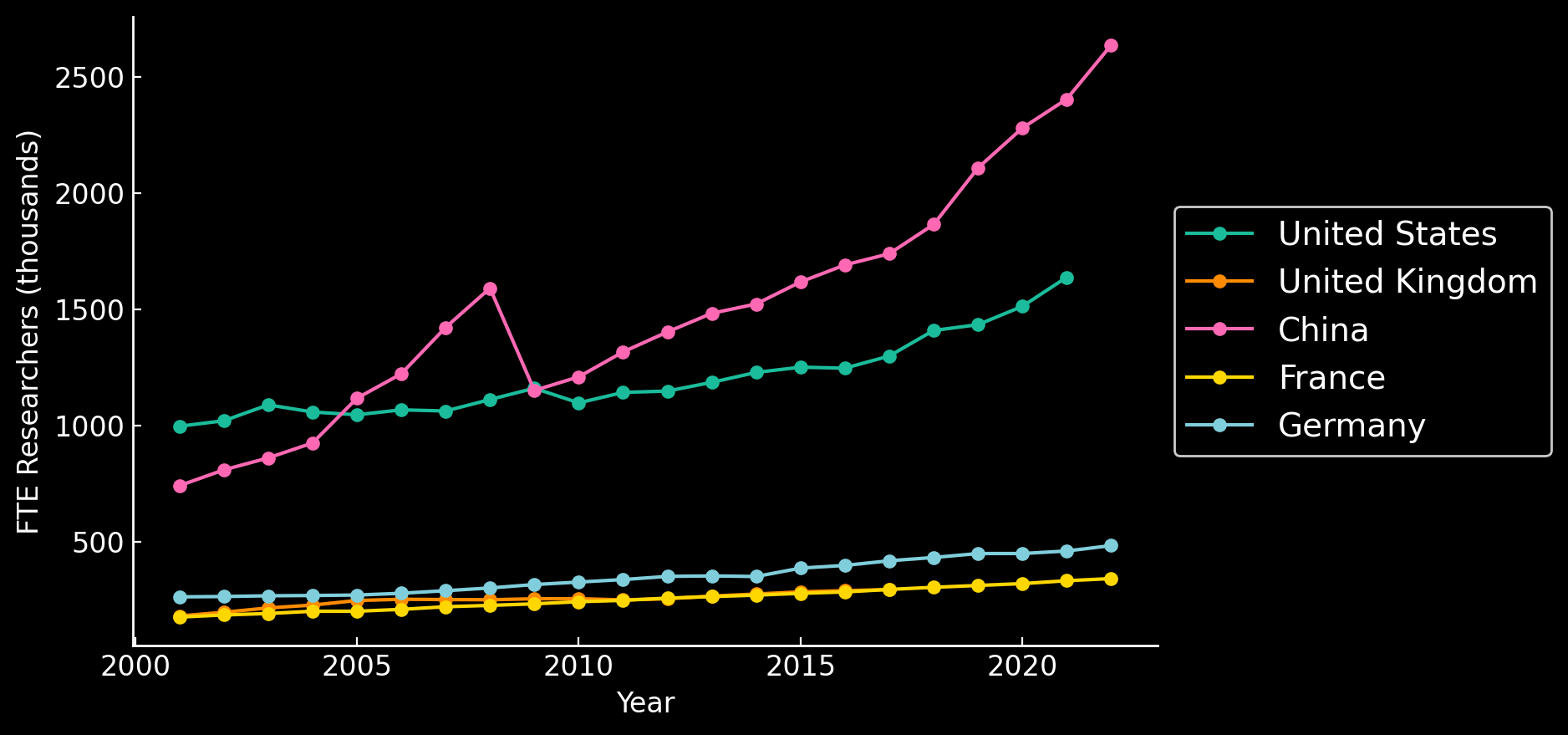
NSF - https://ncses.nsf.gov/pubs/nsf24332; UNESCO UISI SDG9
Intellectual bottlenecks are growing
📝 Increasing paper count ($\approx$15M per year)
🧬 Larger data sets from cheaper
experiments (genome at
$200 per person, $1 / GB of sequencing)
🔍95% decline in disruptive papers since 1980
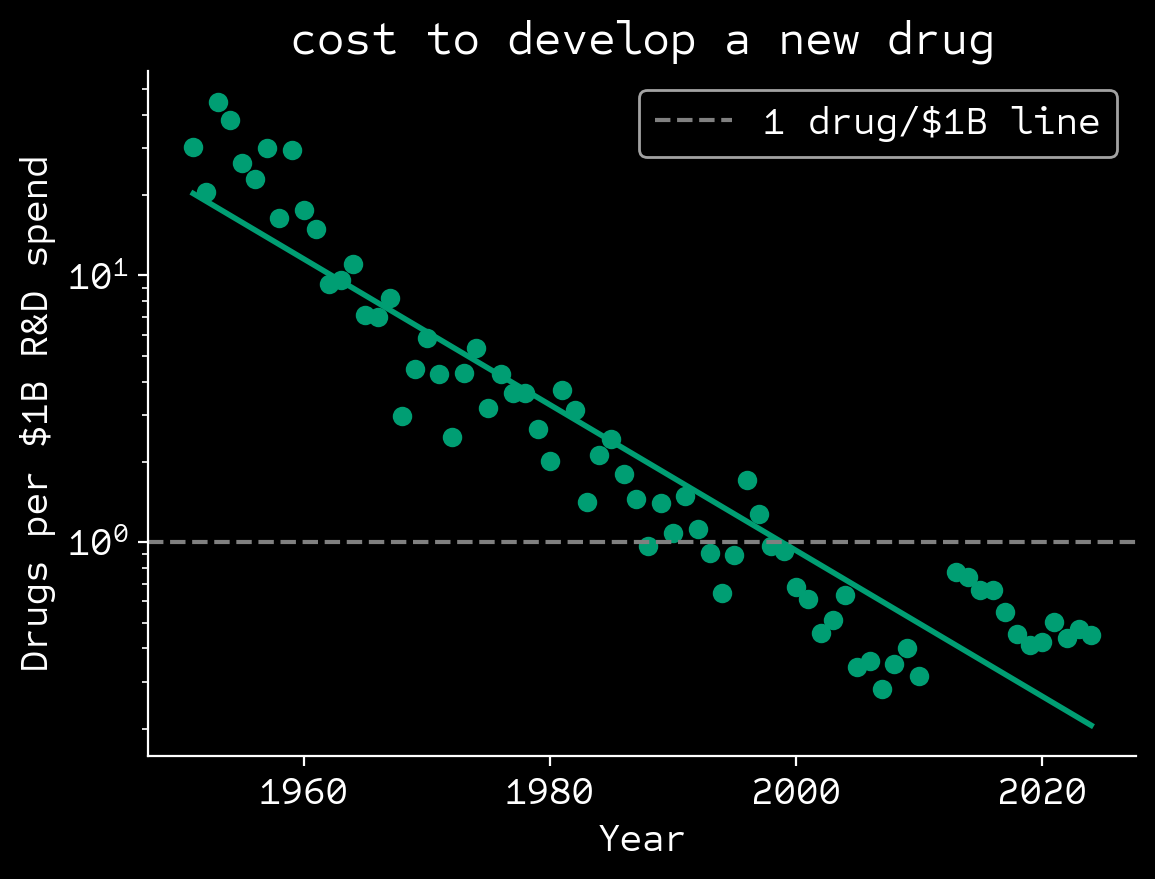
Park, M. et al. Nature 613, 138-144 (2023); Scannell, J.W. et al. Nat. Rev. Drug Discov. 11, 191–200 (2012); Deloitte 2025: Pharma innovation returns.
FutureHouse Mission
Accelerate Scientific Discovery
FutureHouse
- FutureHouse founded in 2023
- Andrew White, Sam Rodriques, Eric Schmidt
- Based in San Francisco with Wetlab
- Raised $35M
FutureHouse Timeline
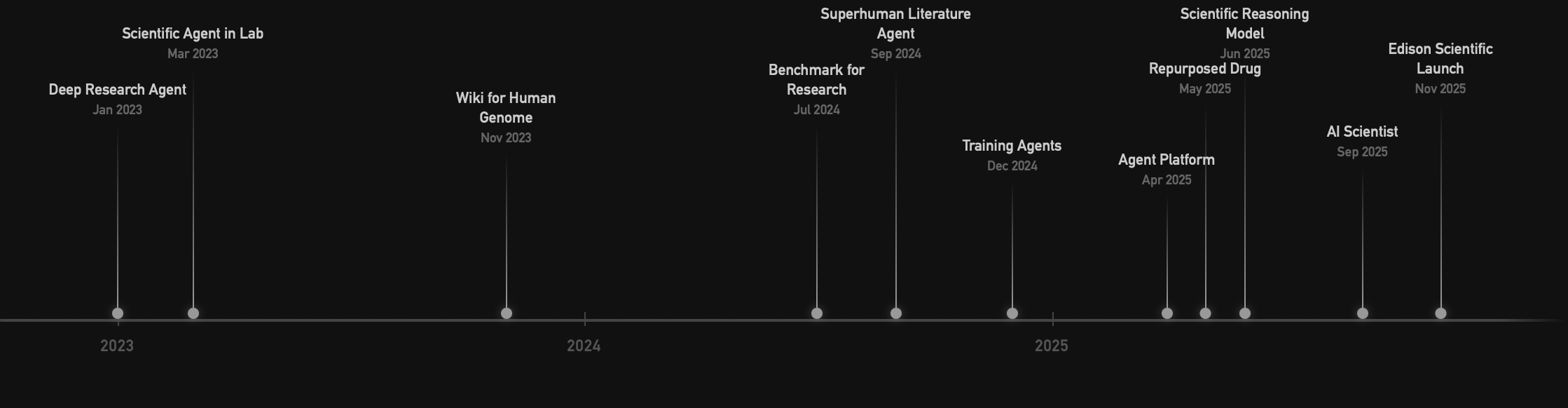
AI Progress
Model intelligence doubles every 7 months
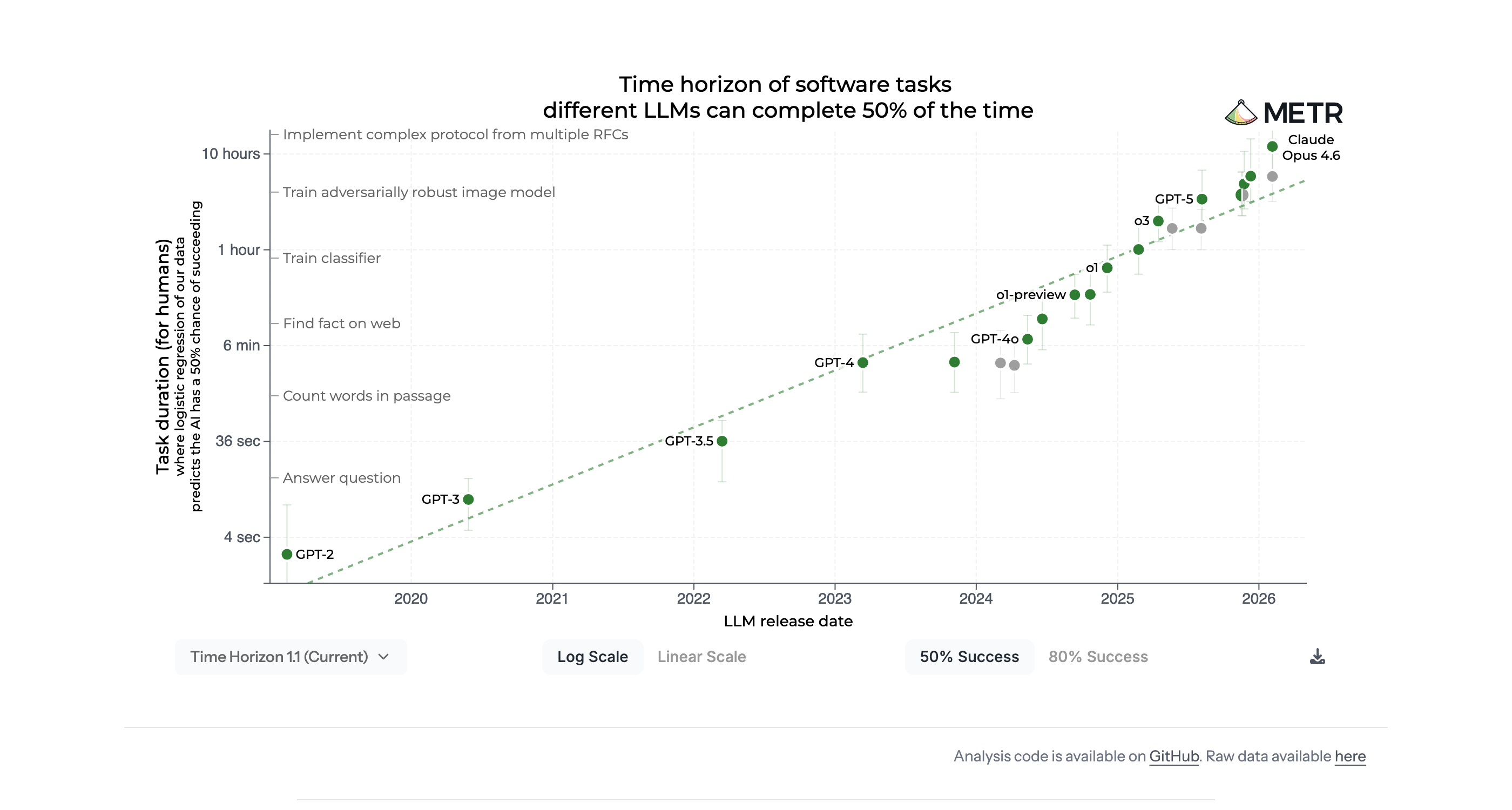
METR Task Completion Benchmark metr.org
Effect is Visible in Economy
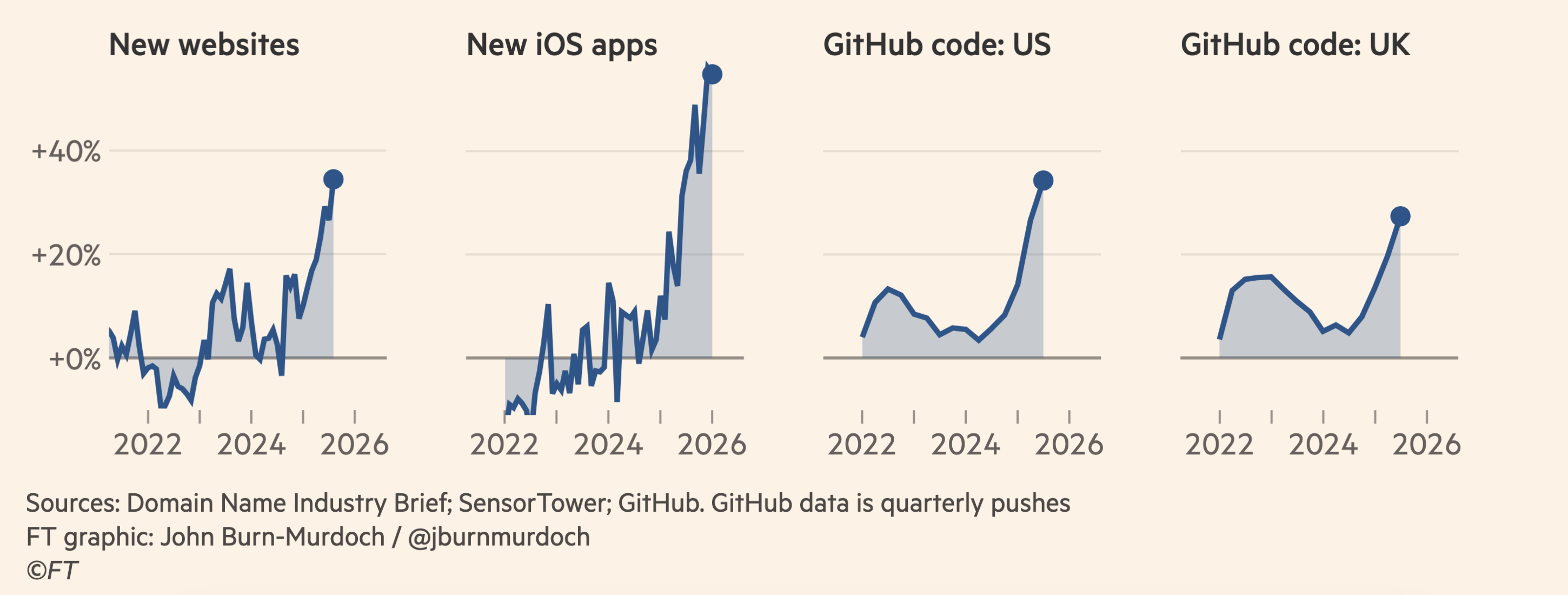
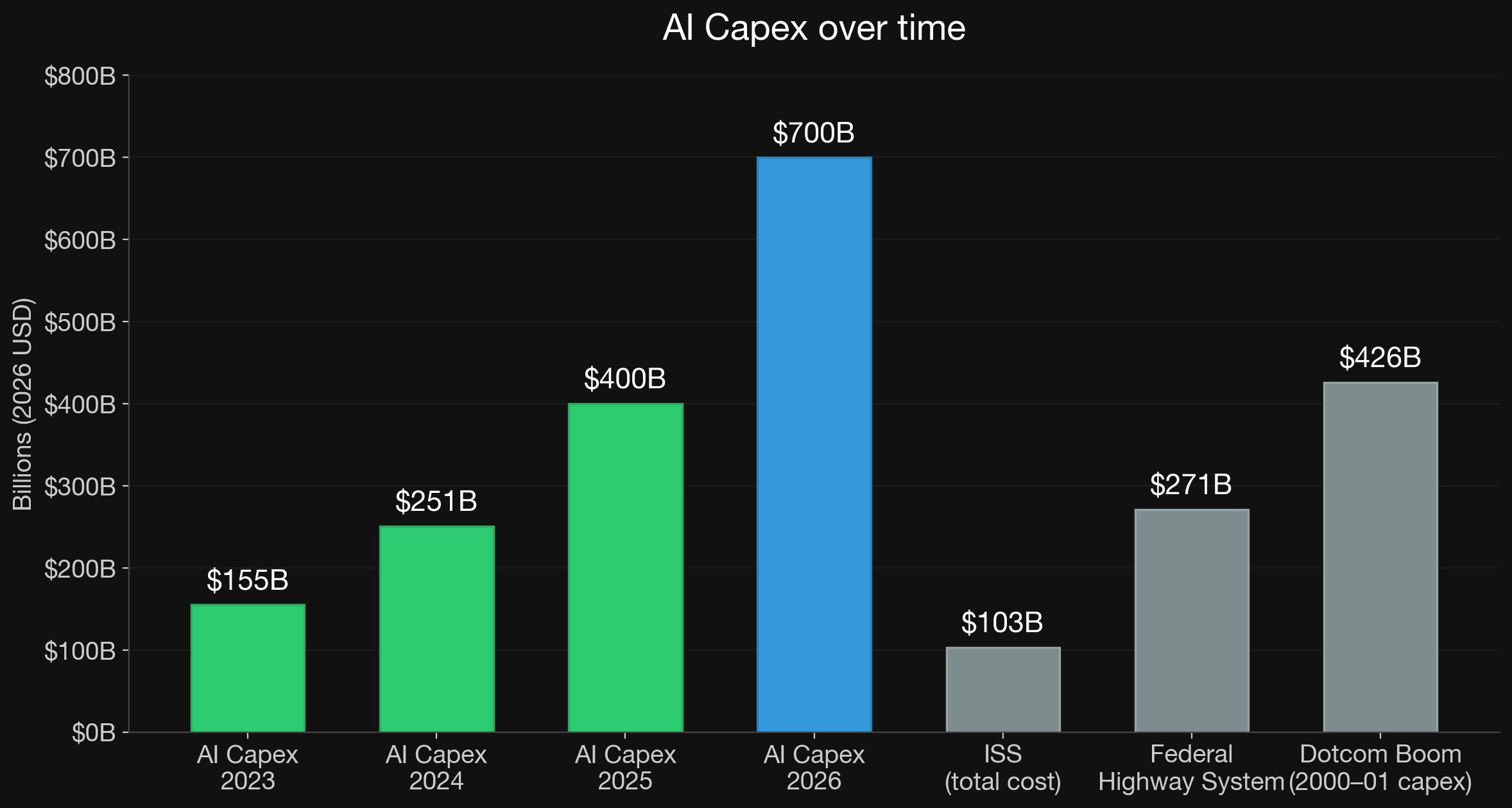
1 NASA OIG Report oig.nasa.gov 2 US FHWA fhwa.dot.gov 3 US Telecom Capex Report ustelecom.org 4 Morgan Stanley AI Market Trends 2026
Science vs Software
- Context: Scientific Literature
- Experiments: Much larger space of actions
- Hypotheses: Hard to choose good hypotheses
Introduction to AI Models
| LLM | model (knowledge, text generation) | Ex: GPT-5 |
| Agent | model + tools + task (can take actions) | Ex: ChemCrow |
| Co-Scientist | agent + conversation + long-running (human-in-the-loop) | Ex: Claude Code |
| AI Scientist | autonomous + long-running + can execute experiments (novel discoveries) | Ex: Kosmos |
What is an agent?
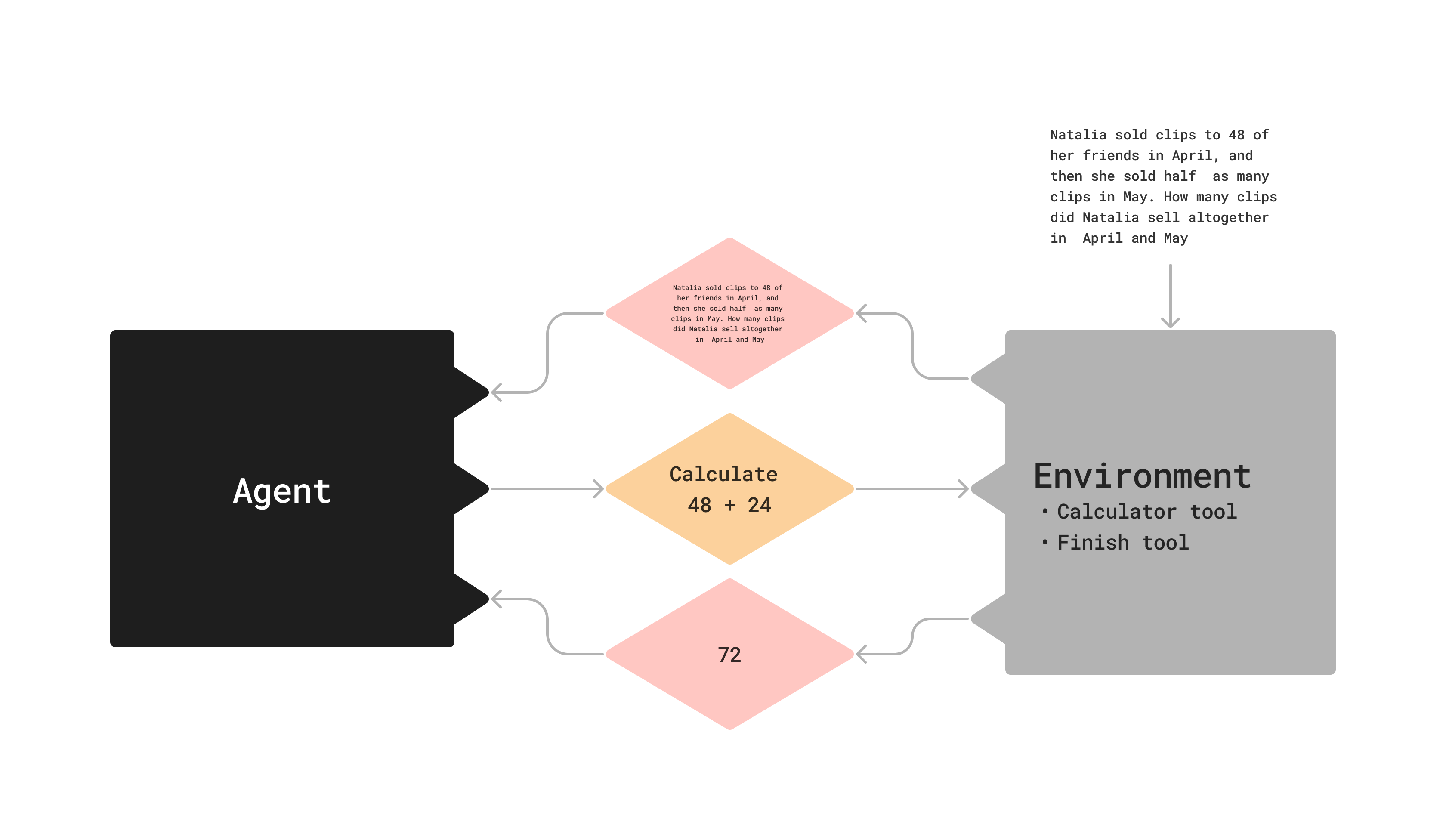
Agent: trained, makes decisions
Environment: untrained, has tools, state
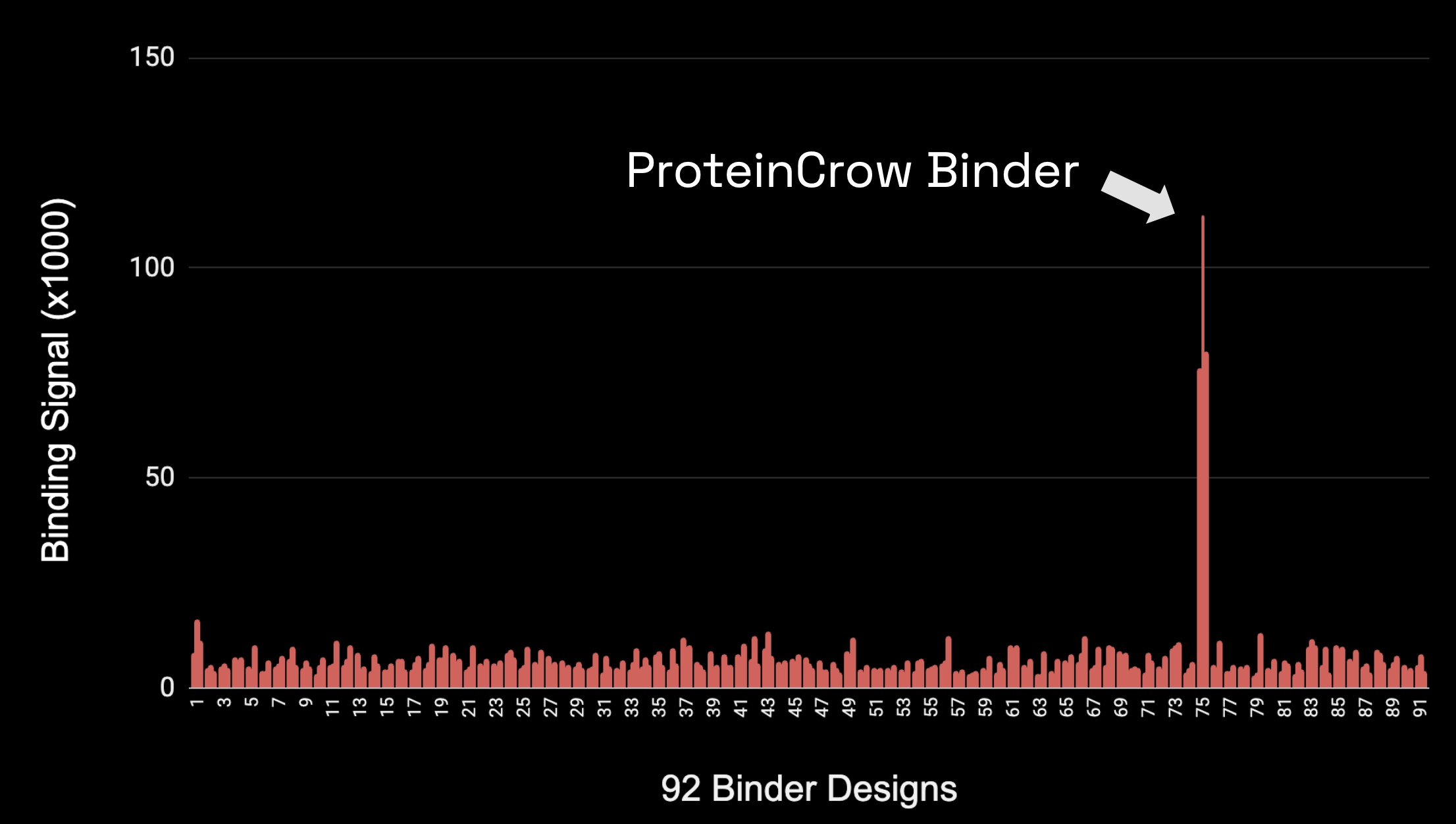
Protein Design Environment
- Protein design with 5 existing deep learning models
- Molecular dynamics, bioinformatics, literature research agent
- Input: "design 92 binders for PD-L1"
Wet lab validation
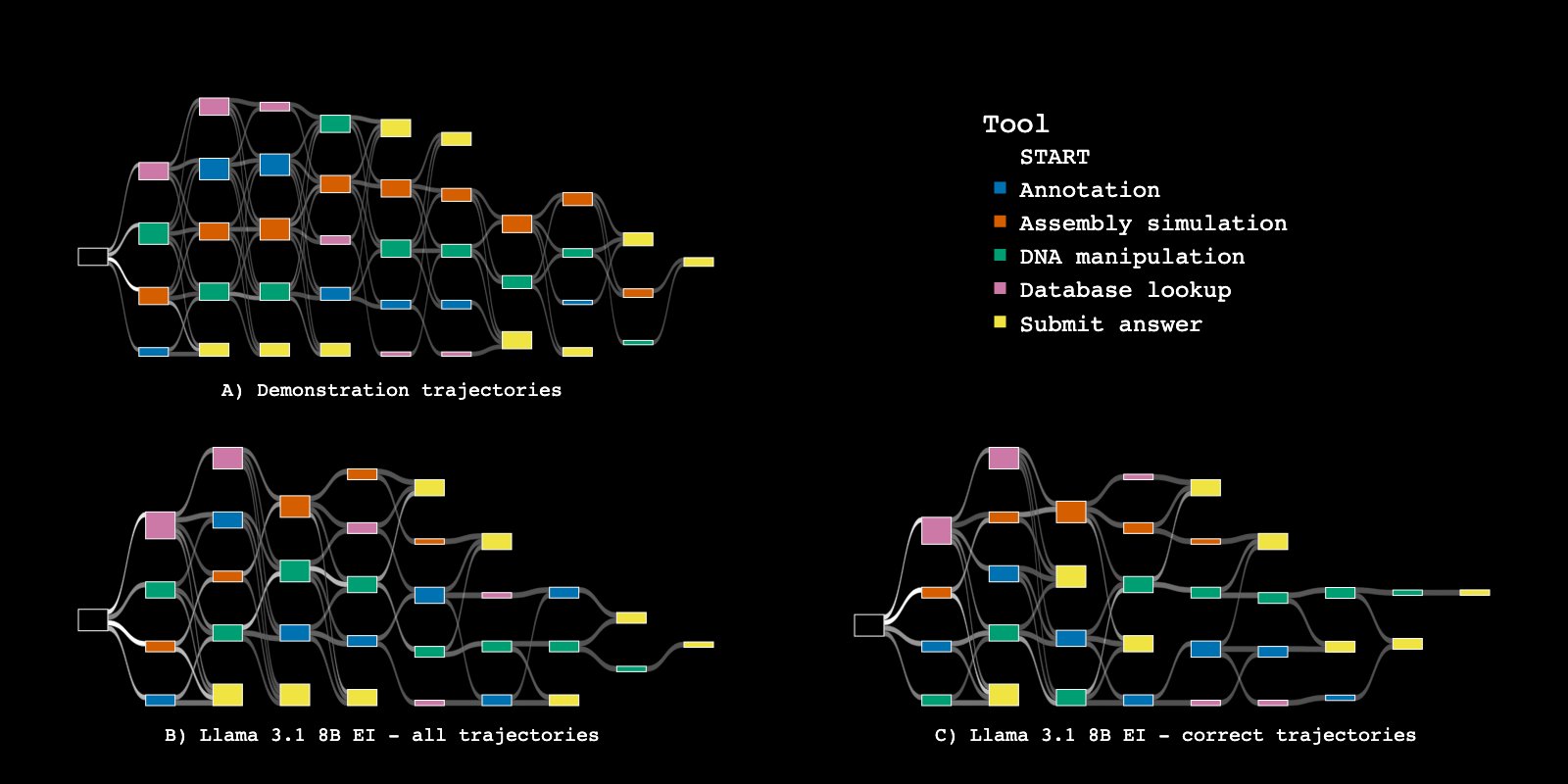
How are agents trained?
- Pre-training — broad knowledge via next-token prediction
- Supervised Fine-Tuning (SFT) — learn from expert instruction–response pairs
- Reinforcement Learning (RL) — optimize through environment interaction with verifiable rewards
RL expands capabilities beyond what supervised data can teach
Verifiable Rewards
Computational checks (code execution, tool outputs) produce objective reward signals — no human feedback needed
LabBench: verifiable tasks as RL training signal
Laurent et al., LabBench2, 2026; Narayanan et al., Aviary, 2024; 2025 NVIDIA Technical Blog
Learning vs Frontier Models
Training Curve
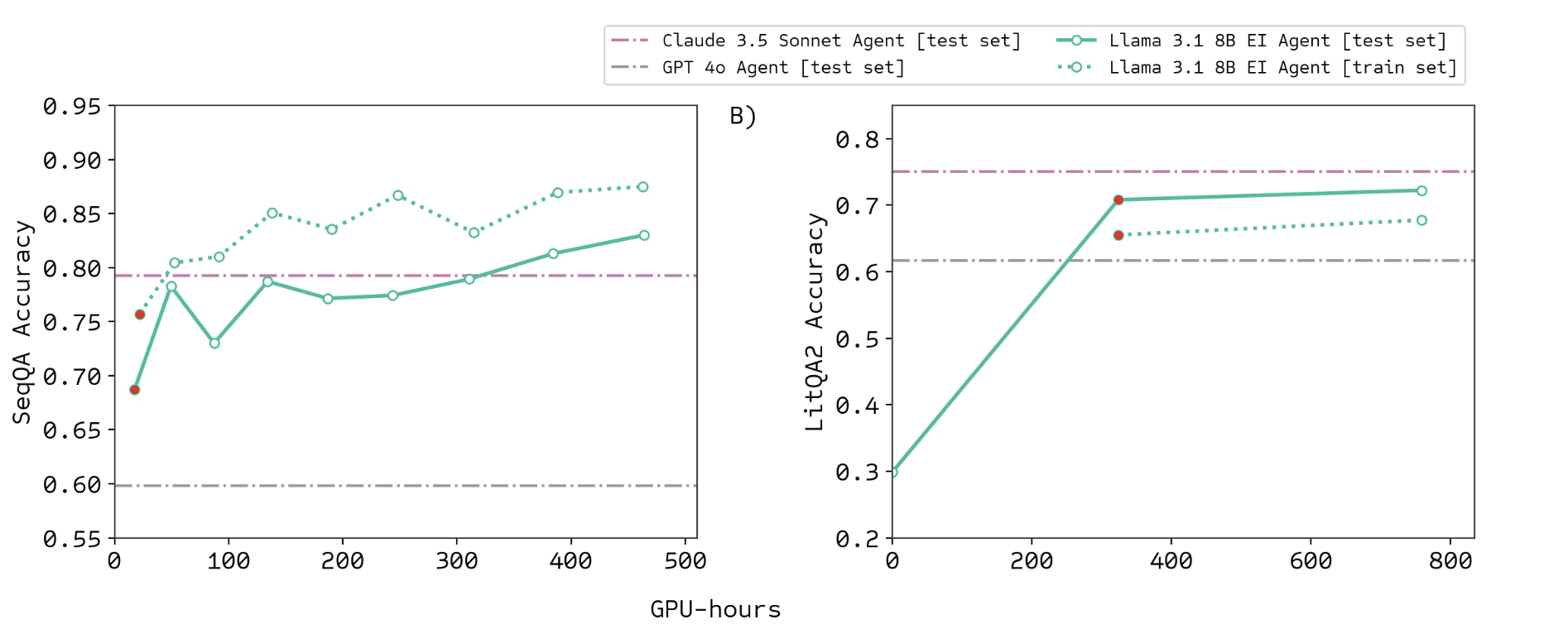
Narayanan et al., Aviary: training language agents on challenging scientific tasks, 2024
FutureHouse Agents
| Name | Environment | Key Tools |
|---|---|---|
| PaperQA | Literature Research | Search, Citation Traversal |
| ProteinCrow | Designing novel proteins | AlphaFold2, Molecular Dynamics |
| ChemCrow/Phoenix | Designing new molecules | Retrosynthesis, self-driving robotic lab |
| Data analysis crow | Generating discoveries from data | bioinformatics tools, code, file system |
Automating research of scientific literature
Language agents achieve superhuman synthesis of scientific knowledge
Michael D. Skarlinski, Sam Cox, Jon M. Laurent, James D. Braza, Michaela Hinks, Michael J. Hammerling, Manvitha Ponnapati, Samuel G. Rodriques, Andrew D. White arXiv:2409.13740, 2024
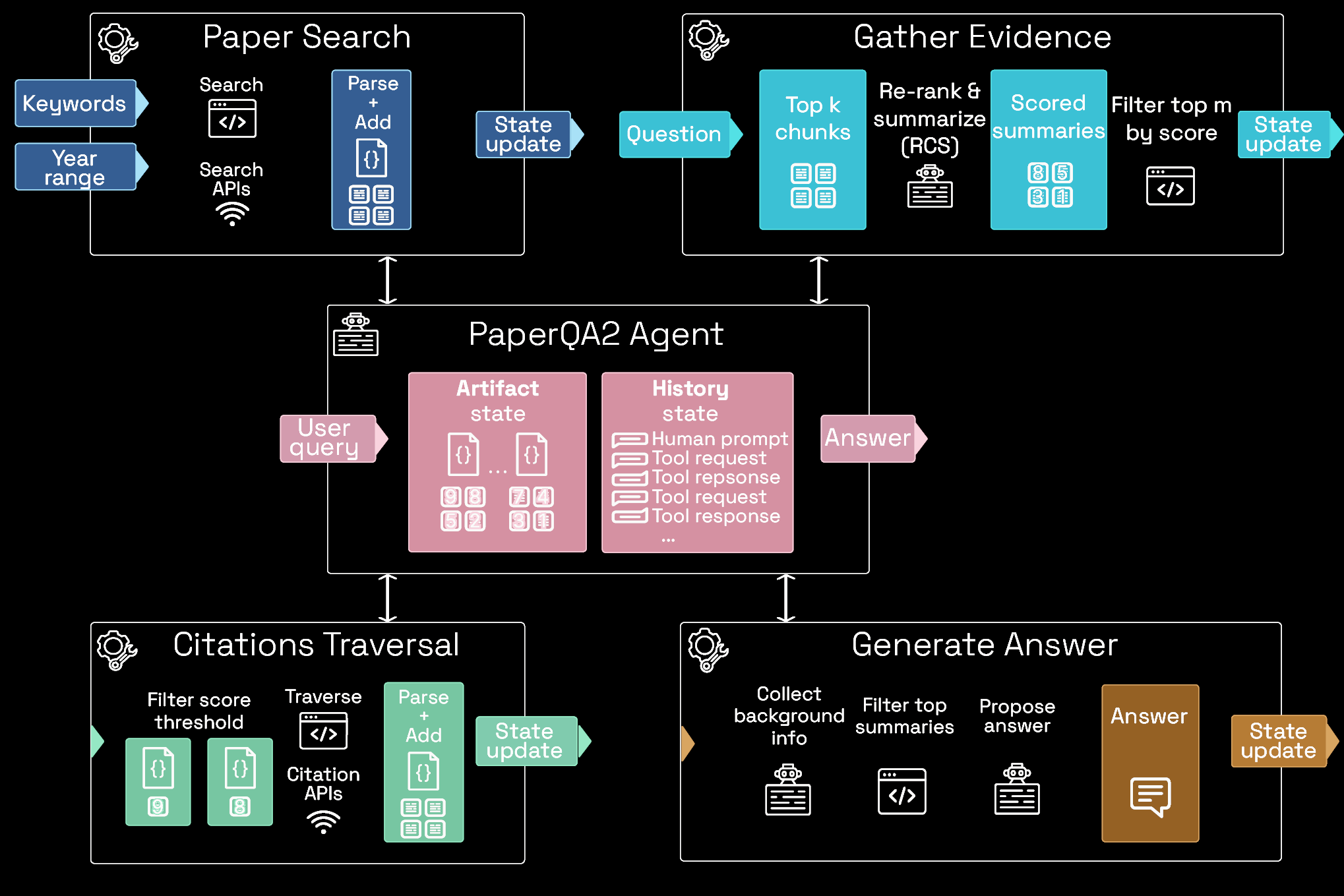
Measurement: LAB-Bench (2024)
Approximately what percentage of Drosophila with a H3.3K36R mutation finish developing and enclose?
- 80%
- 19%
- 50%
- 37%
- 6%
- 94%
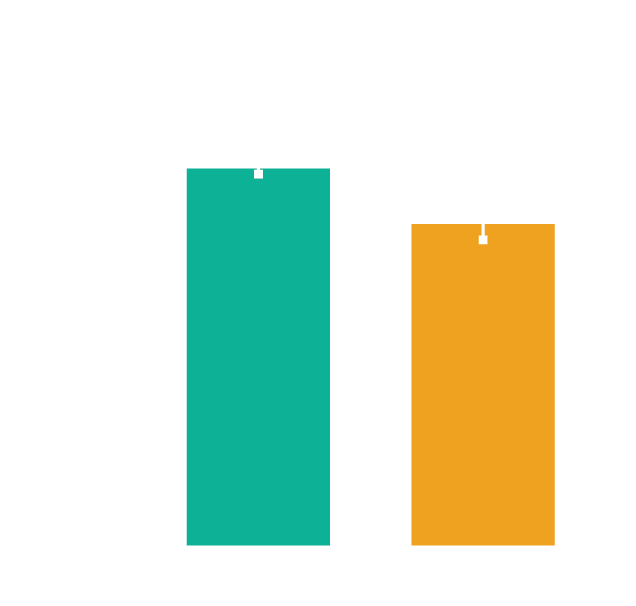
Better at answering questions than PhD biology experts
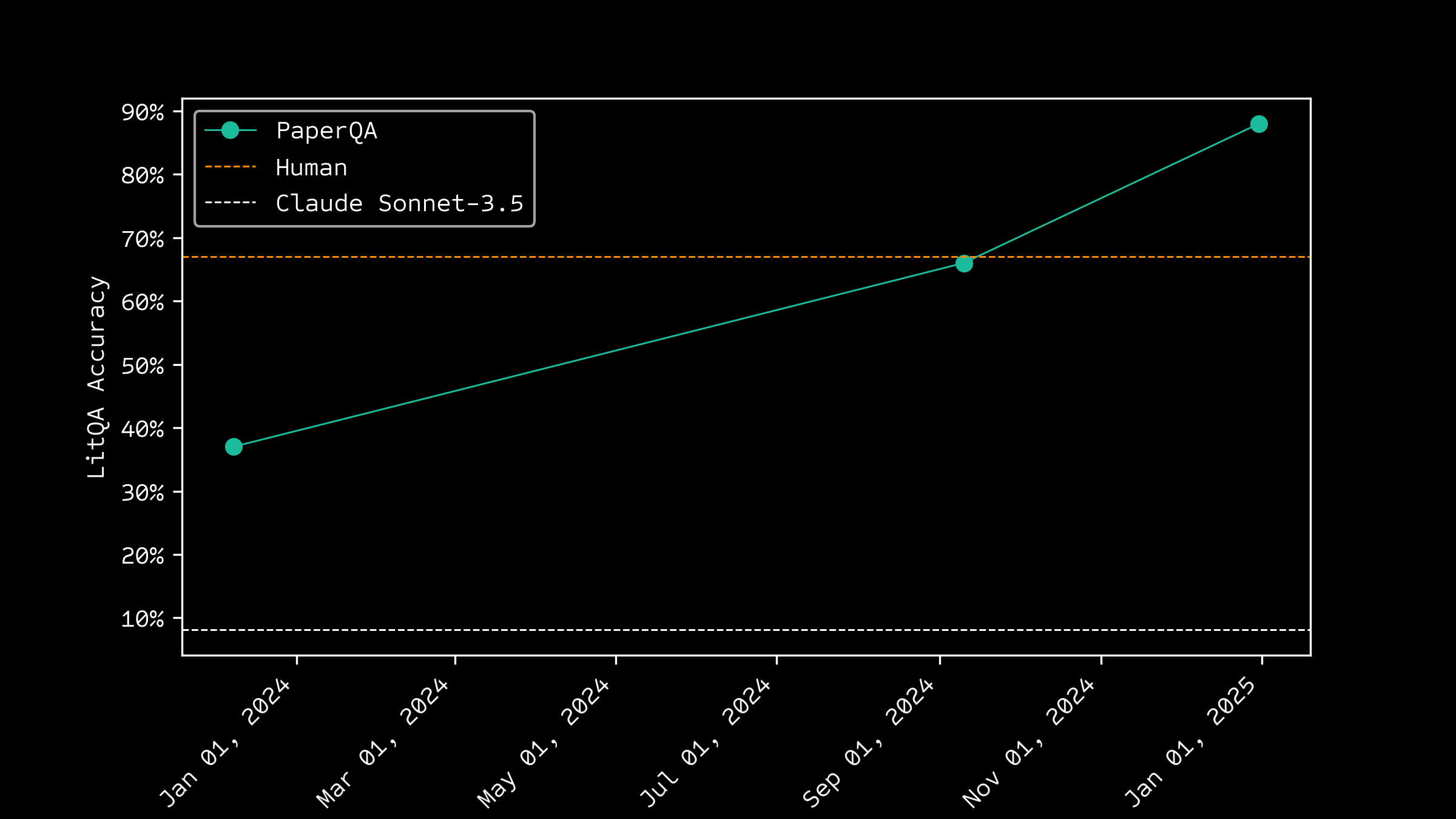
Improving over time
Better than human written Wikipedia articles
PaperQA3 (2026)
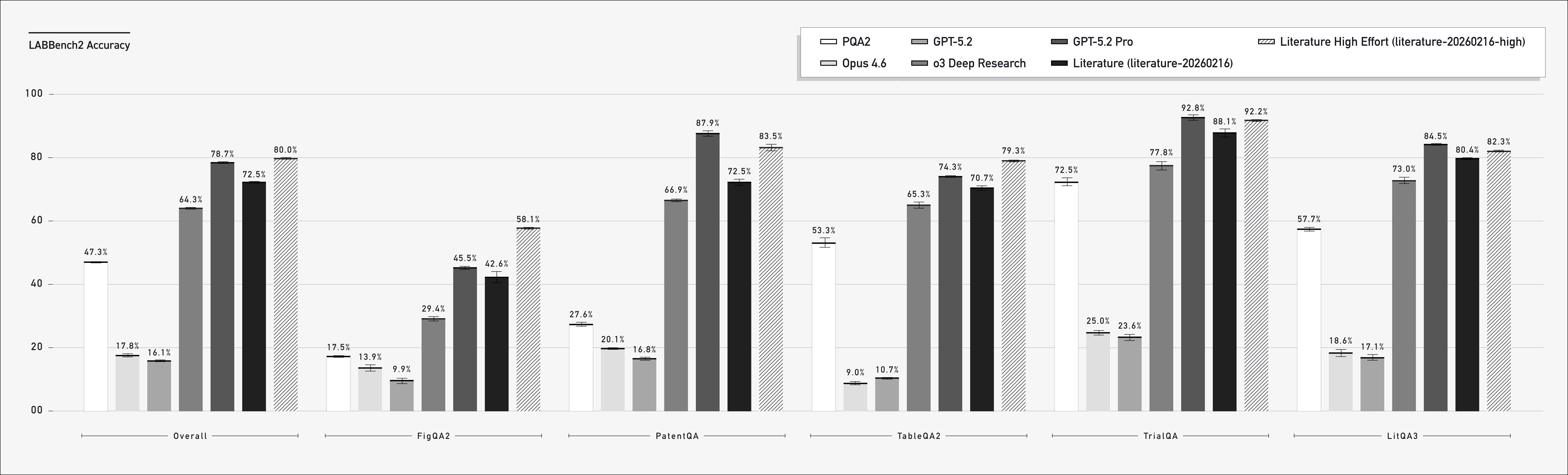
Agents for Data Analysis
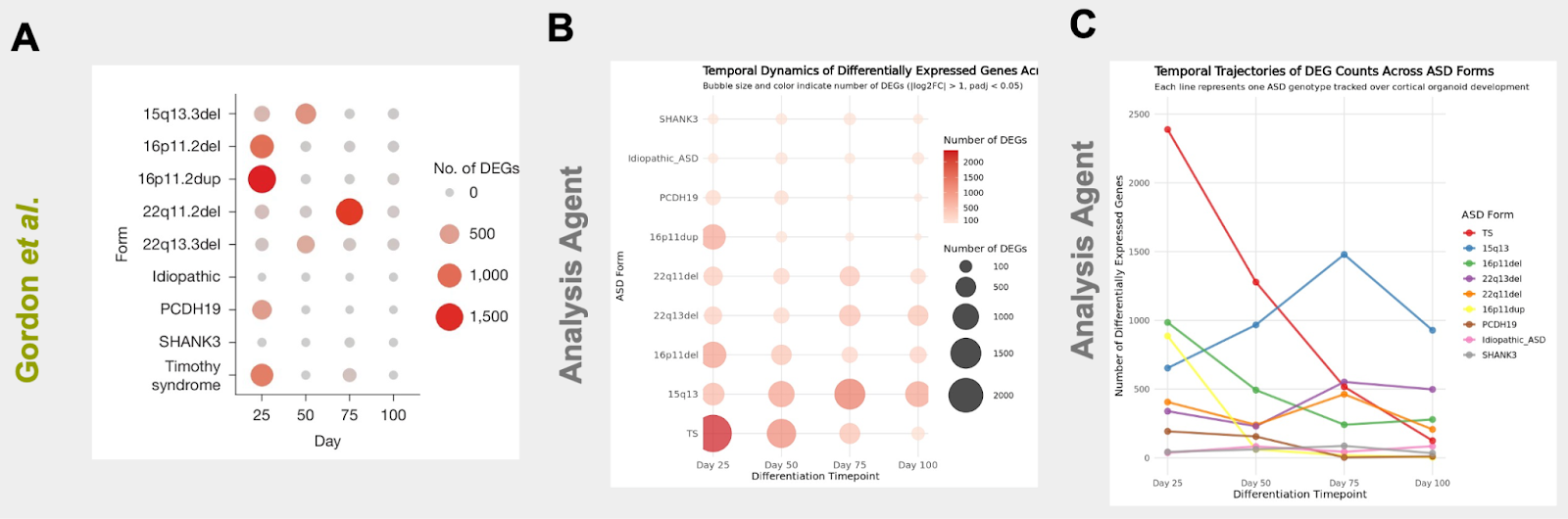
Evaluation: Can it reproduce papers?

... Calculate Spearman correlations of the resulting log-fold change (logFC) values across conditions. Perform hierarchical clustering. Plot and visualize the clustering result as a heatmap to show how different ASD forms cluster together as development progresses
Creates a reproducible R notebook

Side-by-Side
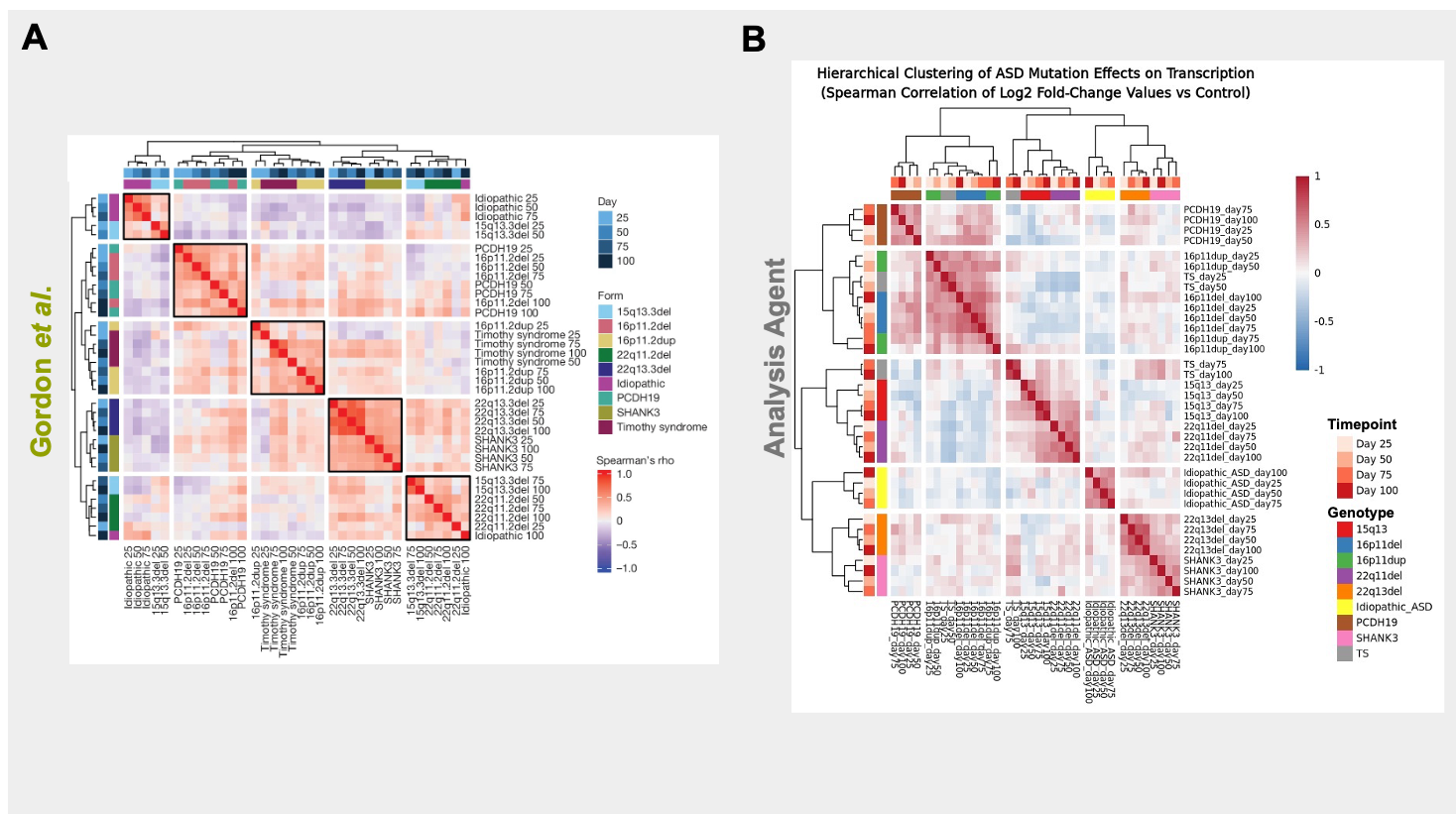
Side-by-Side
BixBench

Mitchener, L., Laurent, J. M., Andonian, A., Tenmann, B., Narayanan, S., Wellawatte, G. P., White, A., Sani, L. & Rodriques, S. G. BixBench: a Comprehensive Benchmark for LLM-based Agents in Computational Biology, 2025.
Can access about 80% of biology data
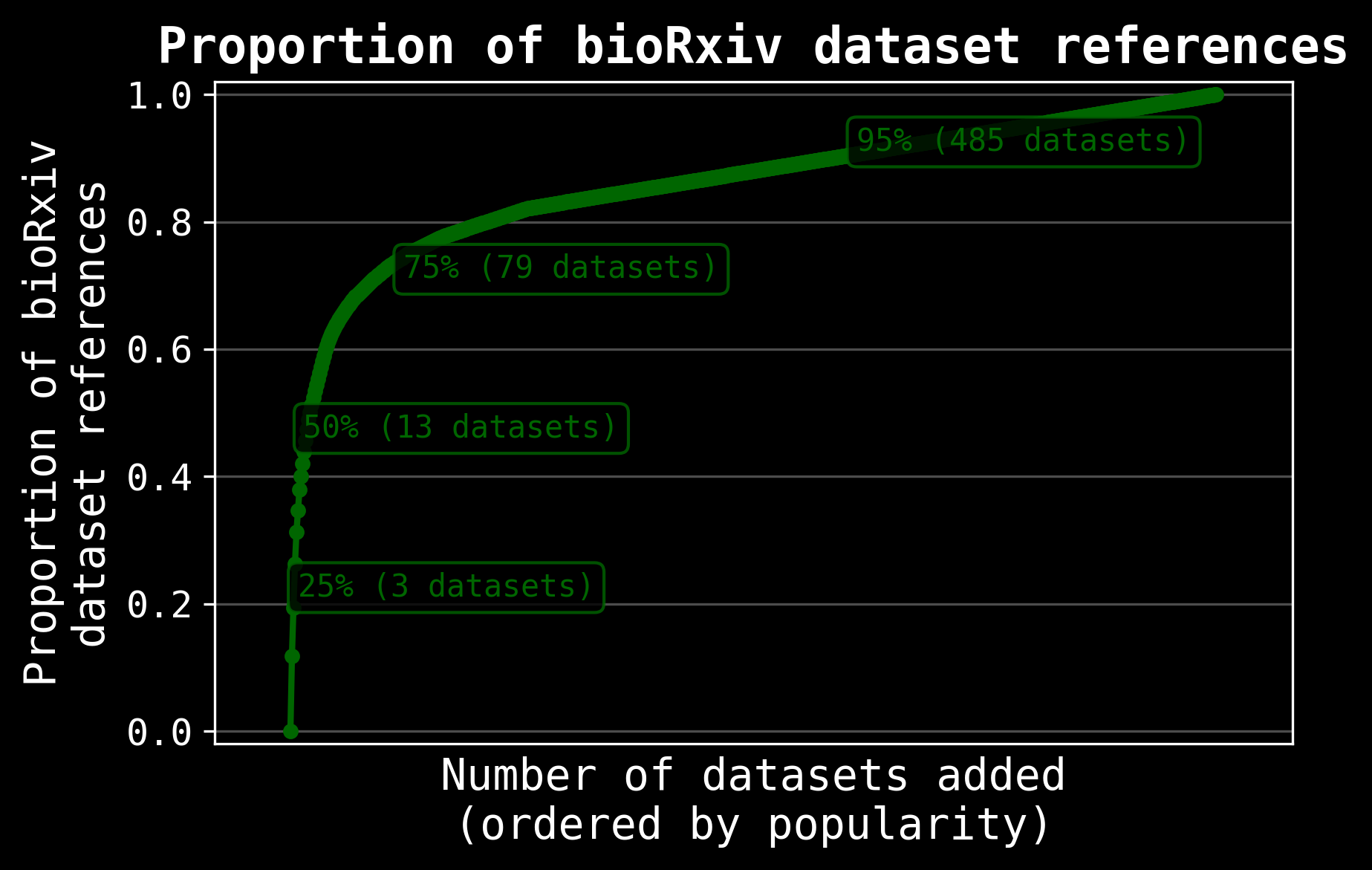
Edison Scientific Platform
- 600 uses per month for academics
- API - can be incorporated into your pipeline/agents
Complete cycle of disease to mechanism to target to drug
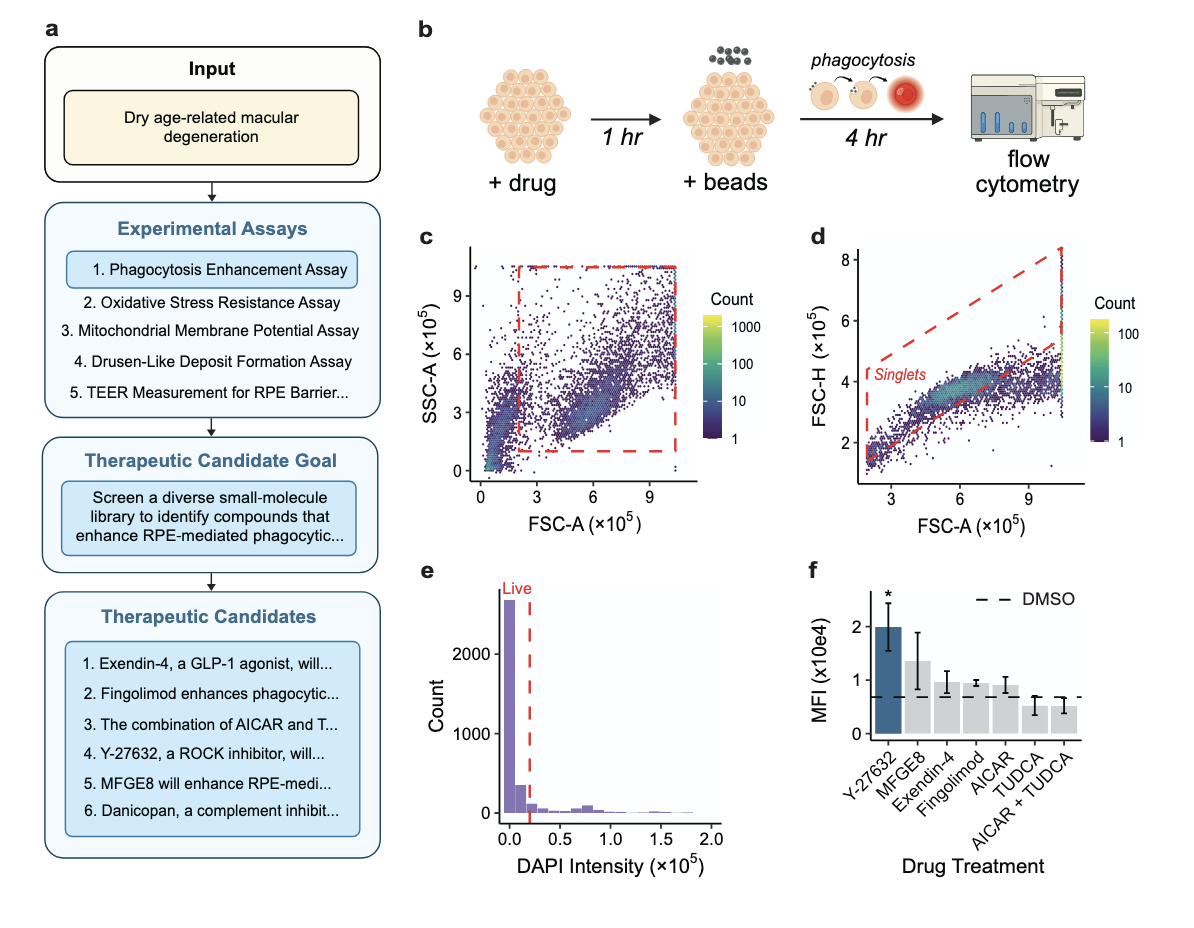
ROBIN: A Multi-Agent System for Automating Scientific Discovery
Ali Essam Ghareeb*, Benjamin Chang*, Ludovico Mitchener, Angela Yiu, Caralyn J. Szostkiewicz, Jon M. Laurent, Muhammed T. Razzak, Andrew D. White†, Michaela M. Hinks‡, Samuel G. Rodriques
Kosmos: An AI Scientist for Autonomous Discovery
Ludovico Mitchener, Angela Yiu, Benjamin Chang, Mathieu Bourdenx, Tyler Nadolski, Arvis Sulovari,
Eric C Landsness, Daniel L Barabasi, Siddharth Narayanan, Nicky Evans, Shriya Reddy, Martha Foiani, Aizad Kamal,
Leah P Shriver, Fang Cao, Asmamaw T Wassie, Jon M Laurent, Edwin Melville-Green, Mayk Caldas, Albert Bou,
Kaleigh F Roberts, Sladjana Zagorac, Timothy C Orr, Miranda E Orr, Kevin J Zwezdaryk, Ali E Ghareeb, Laurie
McCoy, Bruna Gomes, Euan A Ashley, Karen E Duff, Tonio Buonassisi, Tom Rainforth, Randall J Bateman, Michael
Skarlinski, Samuel G Rodriques, Michaela M Hinks, Andrew D White
arXiv:2511.02824, 2025
How do you validate systems like this?
Work with external groups. Input is their experimental data. Three discoveries reproduced in unpublished work. Four novel discoveries.

Entorhinal Cortex
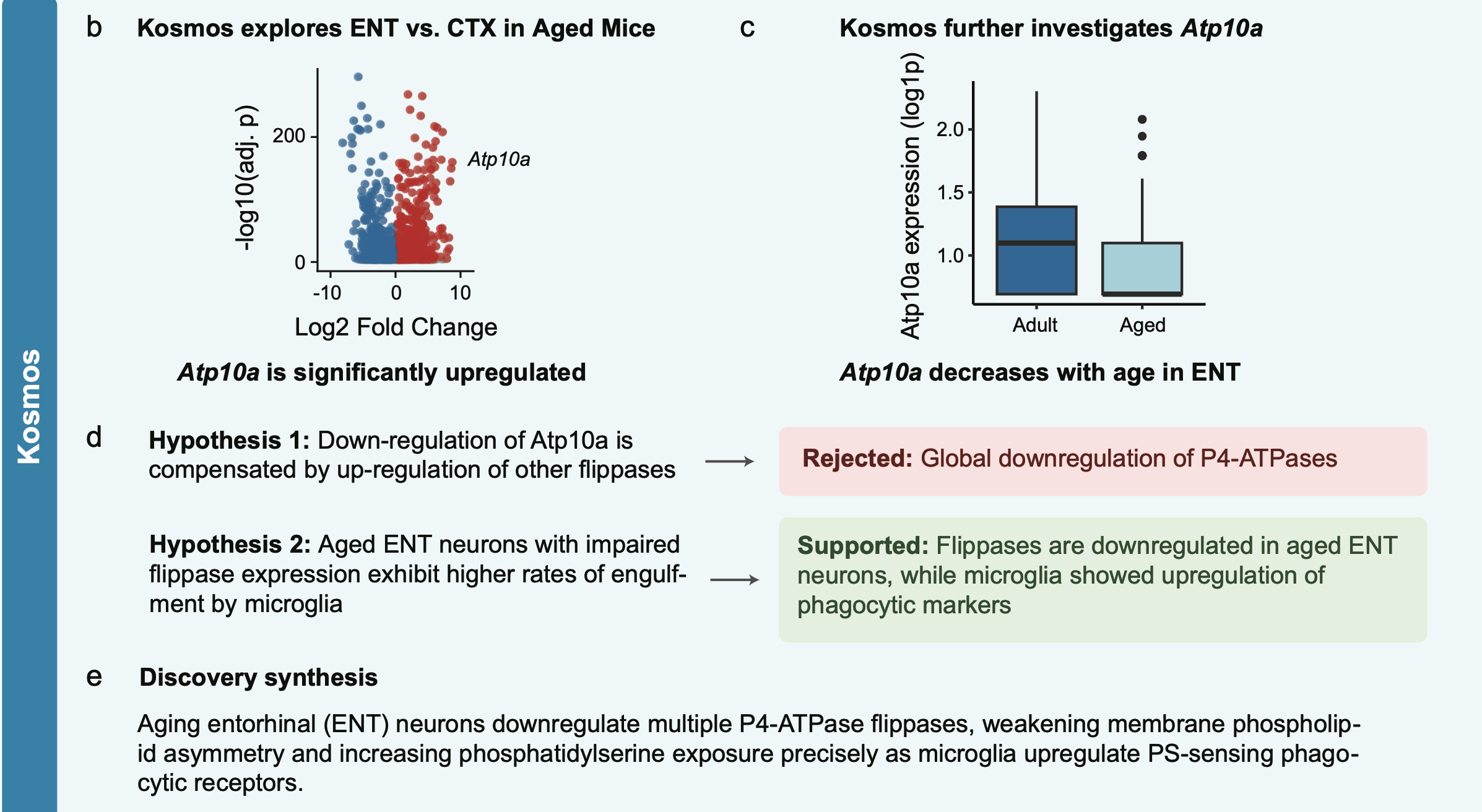
Kosmos overview
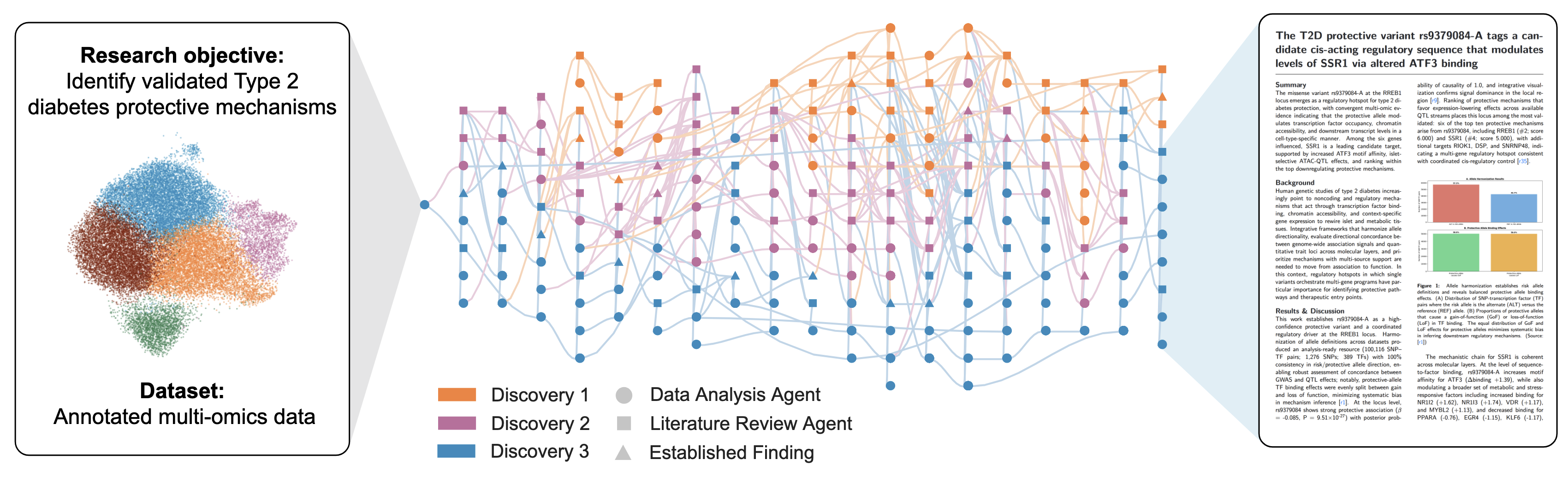
independent expert annotation of task difficulty and correctness
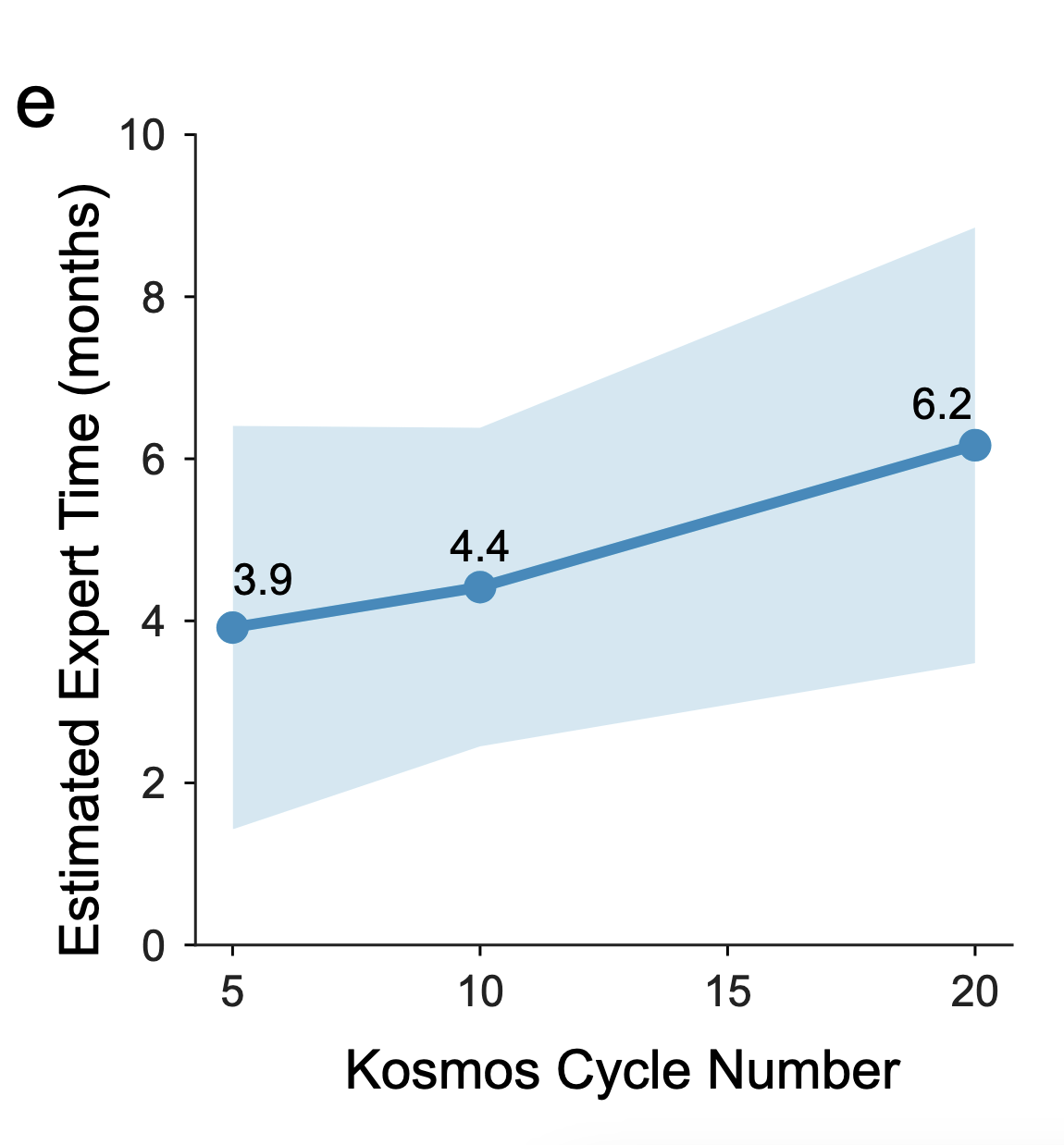
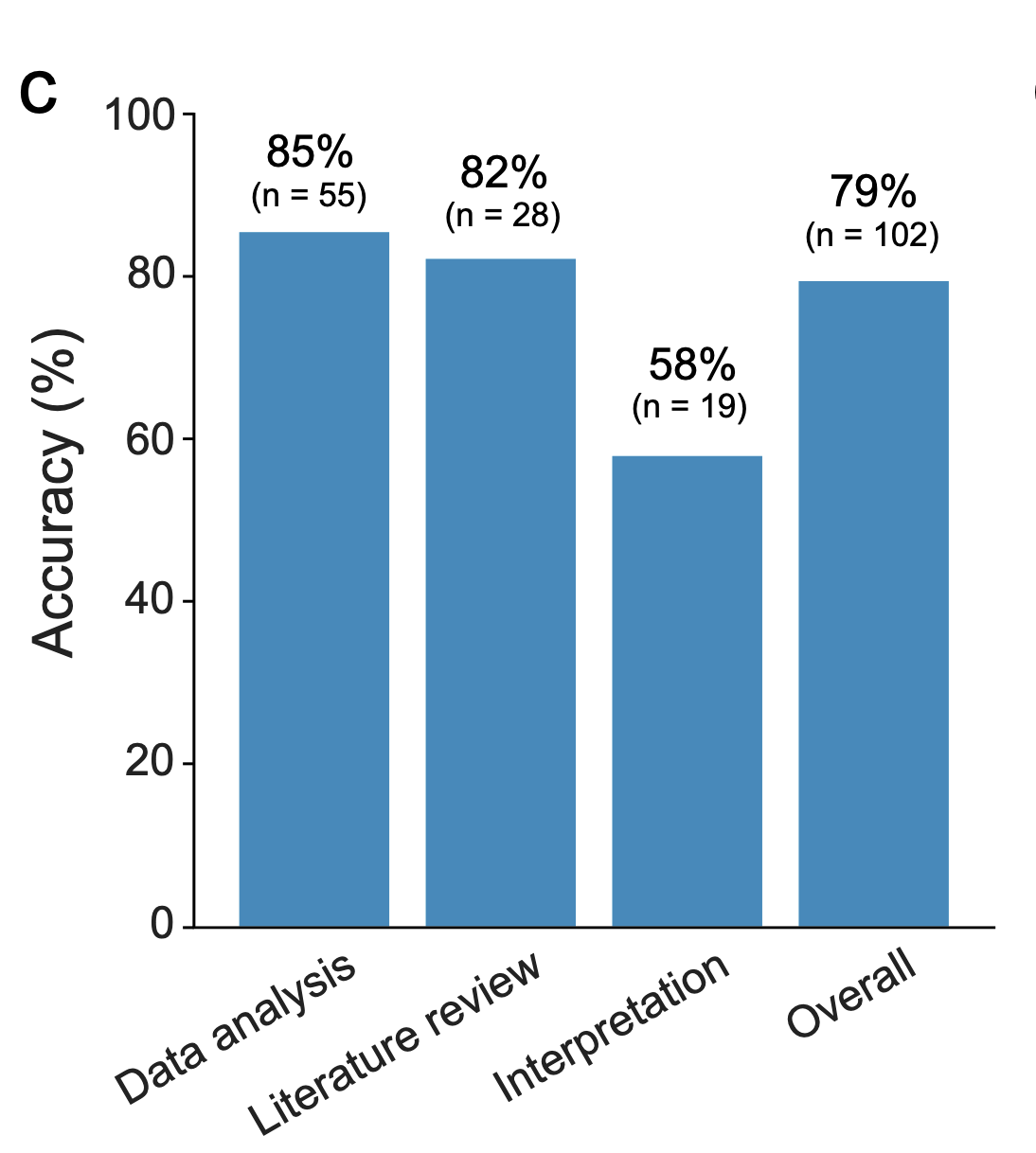
FutureHouse Timeline

Edison Scientific
- Spinout from FutureHouse formed in 2025
- AIxBio Research Lab
- 50 employees
- $70M in seed financing
Scientific Reasoning Models
Training a Scientific Reasoning Model for Chemistry
Siddharth M. Narayanan, James D. Braza, Ryan-Rhys Griffiths, Albert Bou, Geemi Wellawatte, Mayk Caldas Ramos, Ludovico Mitchener, Samuel G. Rodriques, Andrew D. White NeurIPS, 2025
Improving Models
| Pretraining | Large Data, Large Compute |
| Scaffolding | Domain knowledge |
| RL with verifiable rewards | Domain knowledge, small data, small compute |
Reasoning scaling
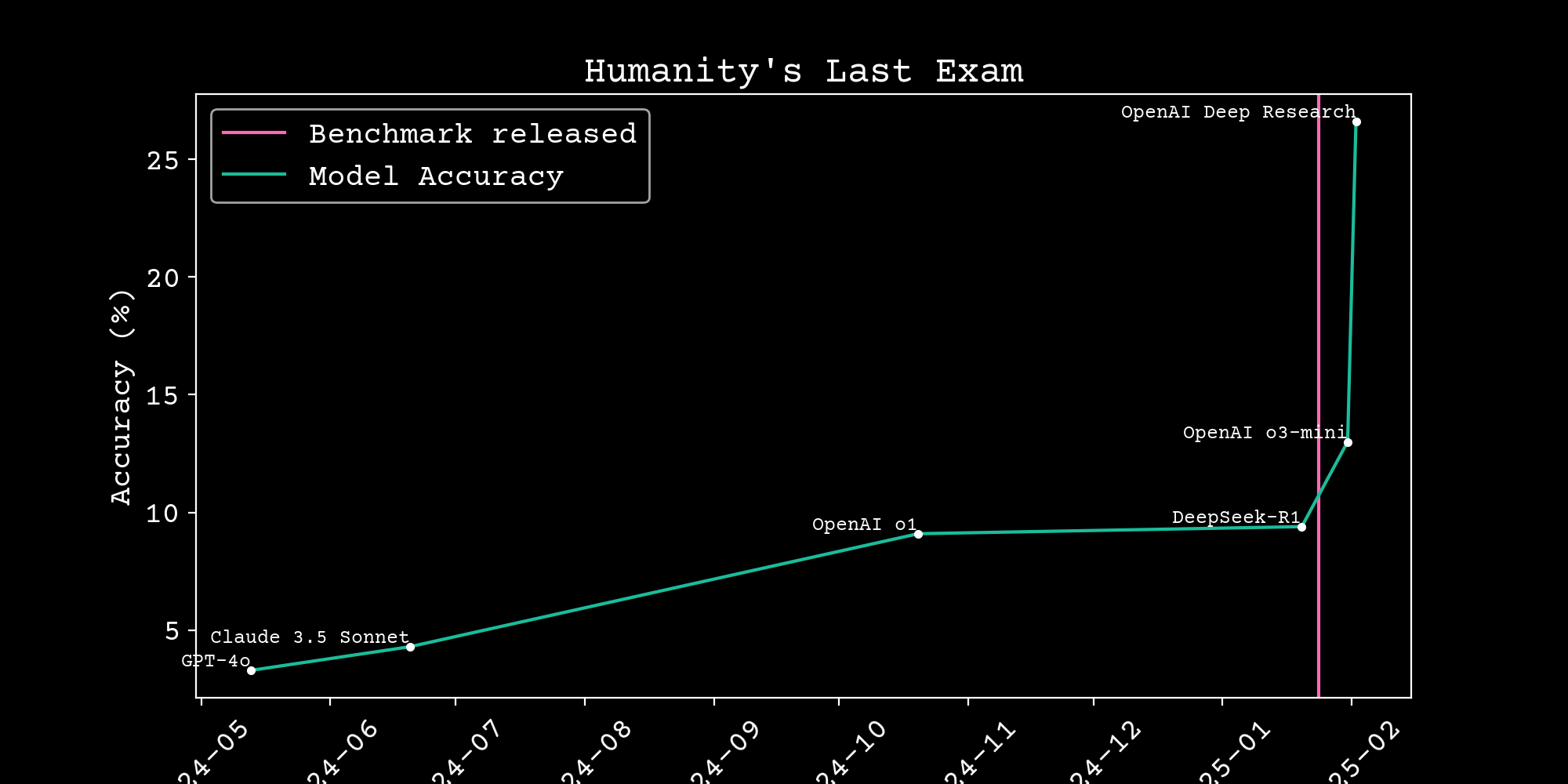
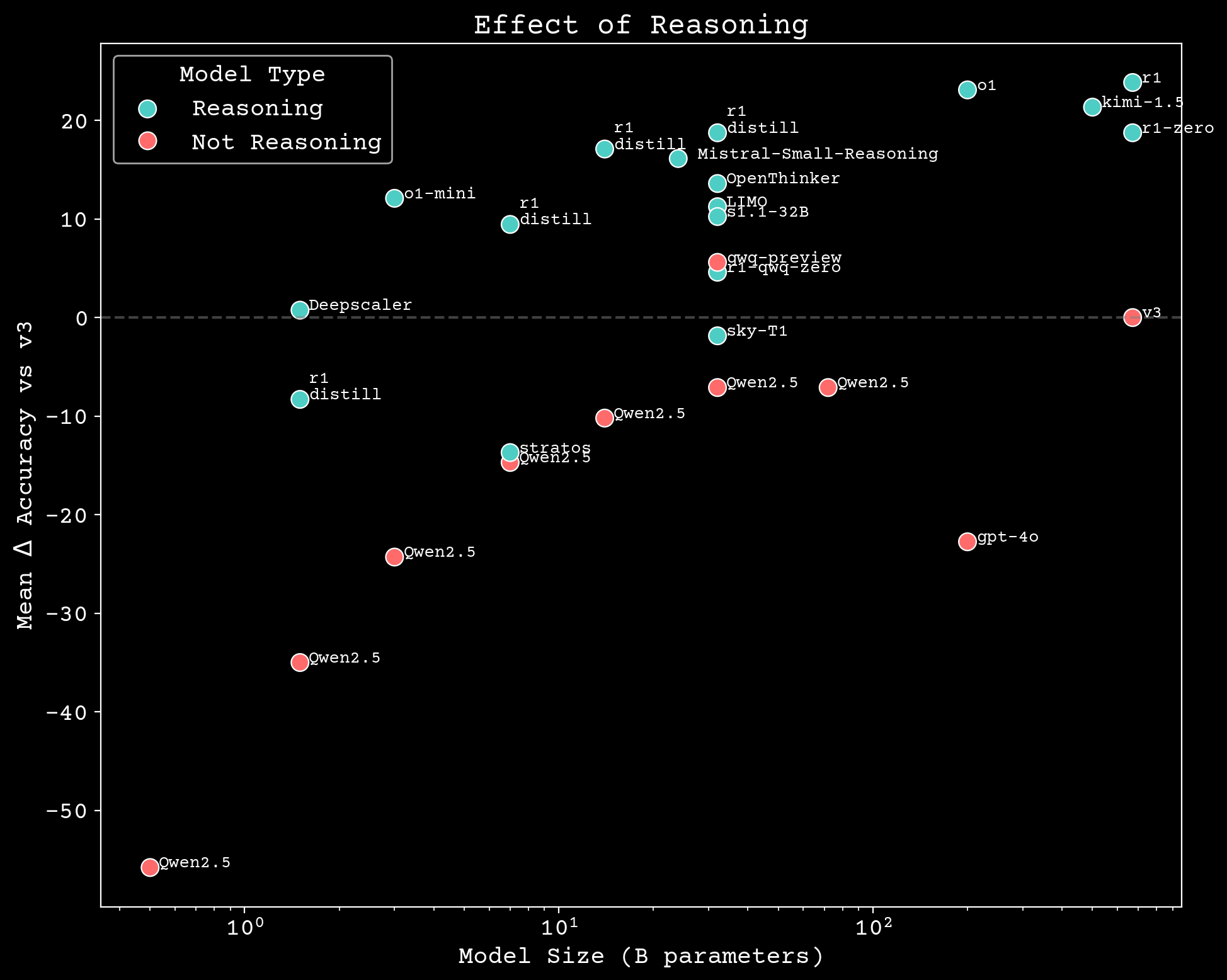
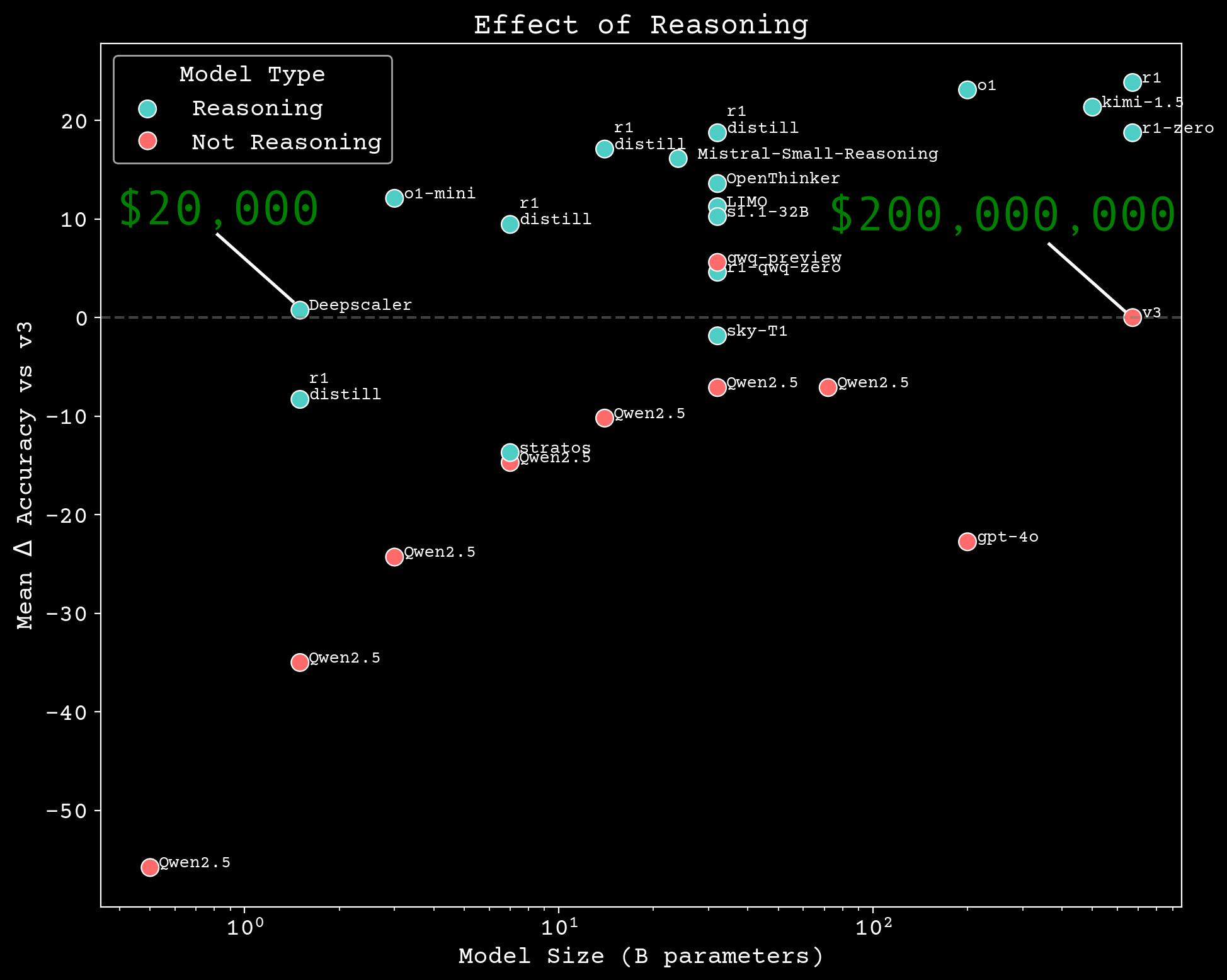
Can we build scientific reasoning models?
chemistry reasoning model
Works with molecular structures, but reasons in English
Start from base LLM and teach it chemistry
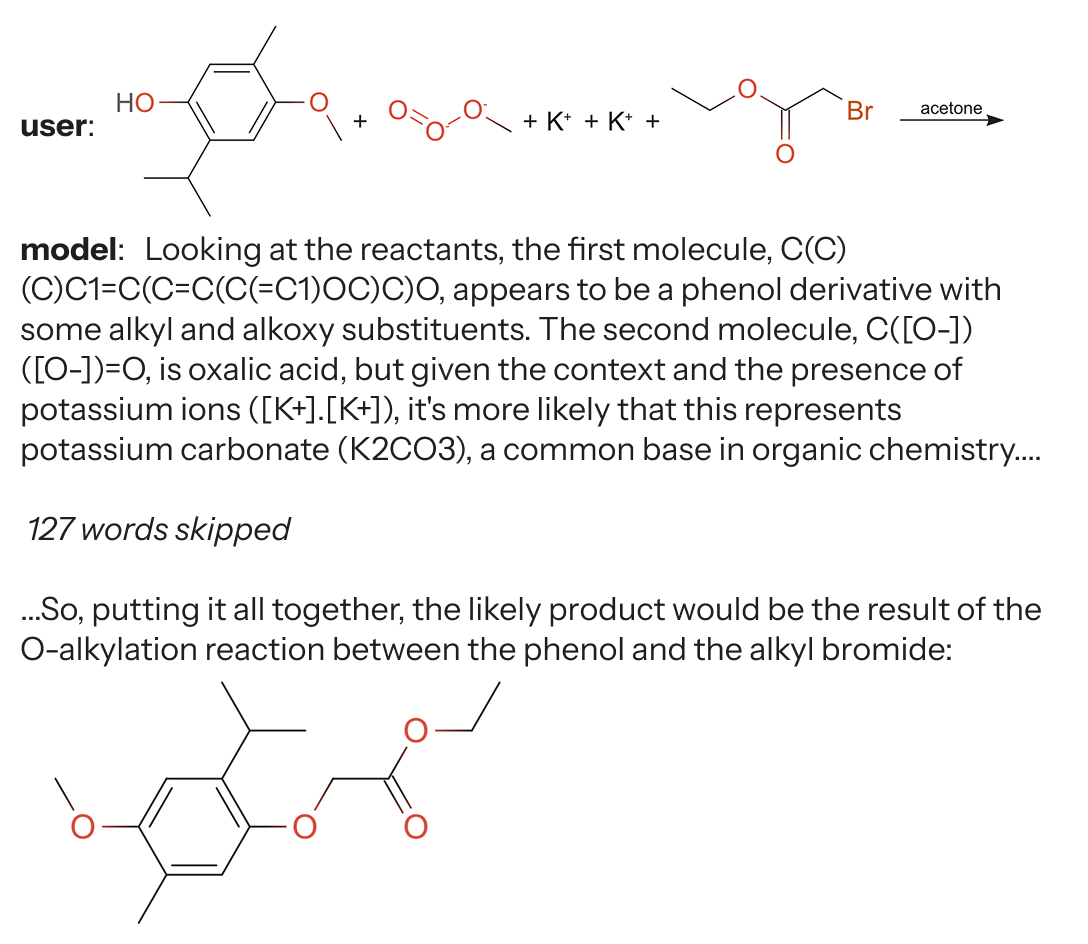
What can a reasoning model do?
Q:Propose a 1-step synthesis path that uses only commercially available reagents
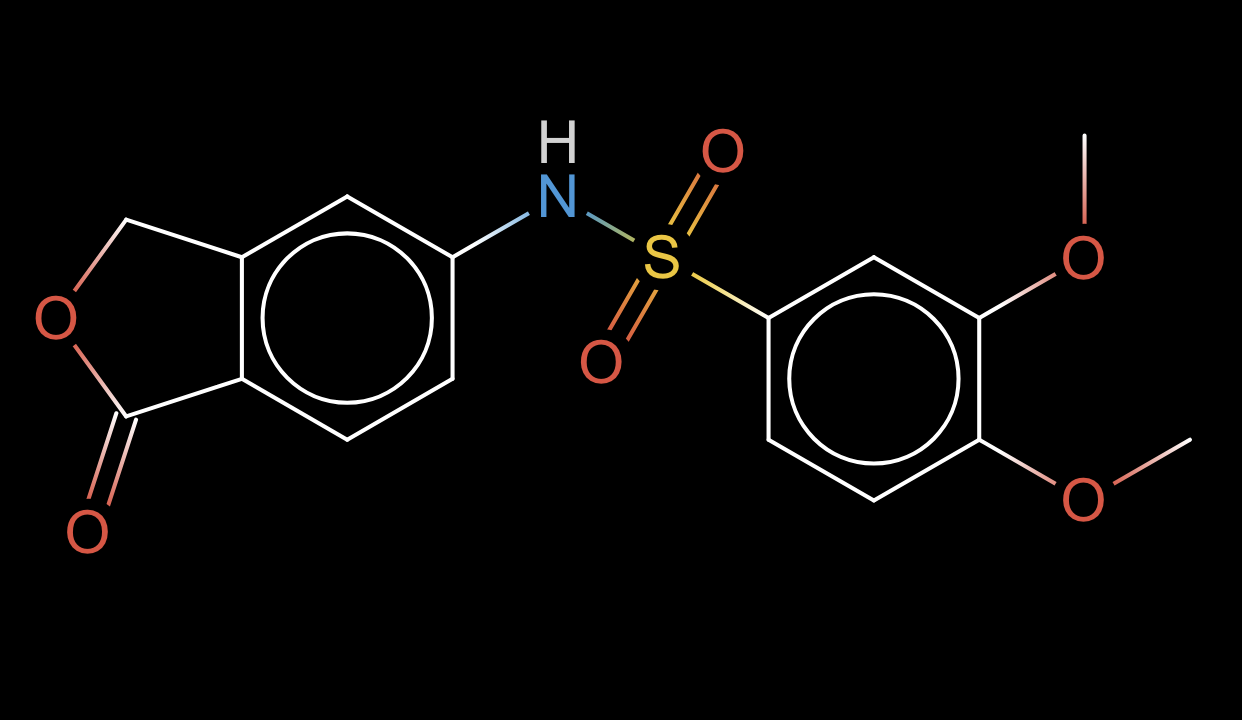
Q: Propose a modification to this molecule to increase its solubility by about 1 LogS unit without affecting its scaffold.
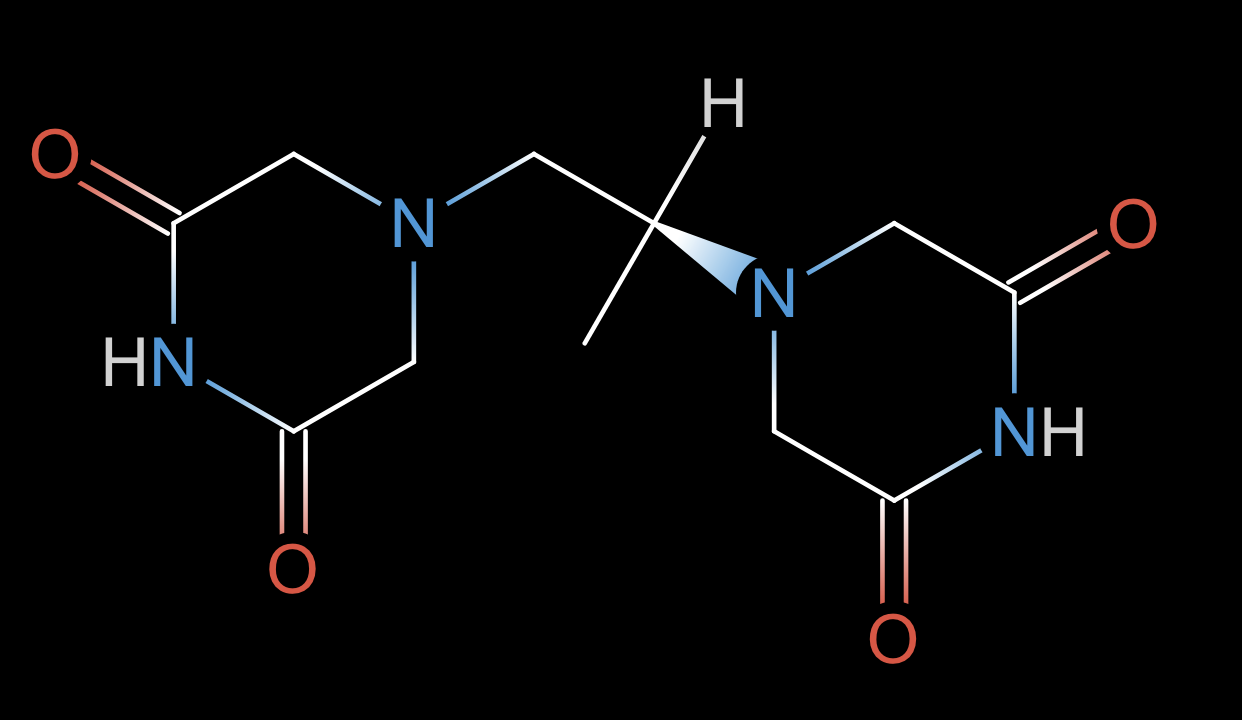
data
| Task | Subtasks | Examples | Verifier | Templates | Data source name |
|---|---|---|---|---|---|
| functional group | 1 | 74562 | code | 6 | ChEMBL |
| organism molecular formula | 1 | 74164 | molecule comparison | 10 | COCONUT |
| IUPAC name | 1 | 74994 | code | 10 | COCONUT |
| SMILES completion | 1 | 74990 | code | 10 | COCONUT |
| solubility edit | 3 | 115977 | ML model, code | 15 | ChEMBL |
| scent | 180 | 4240 | multiple choice | 8 | pyFUME |
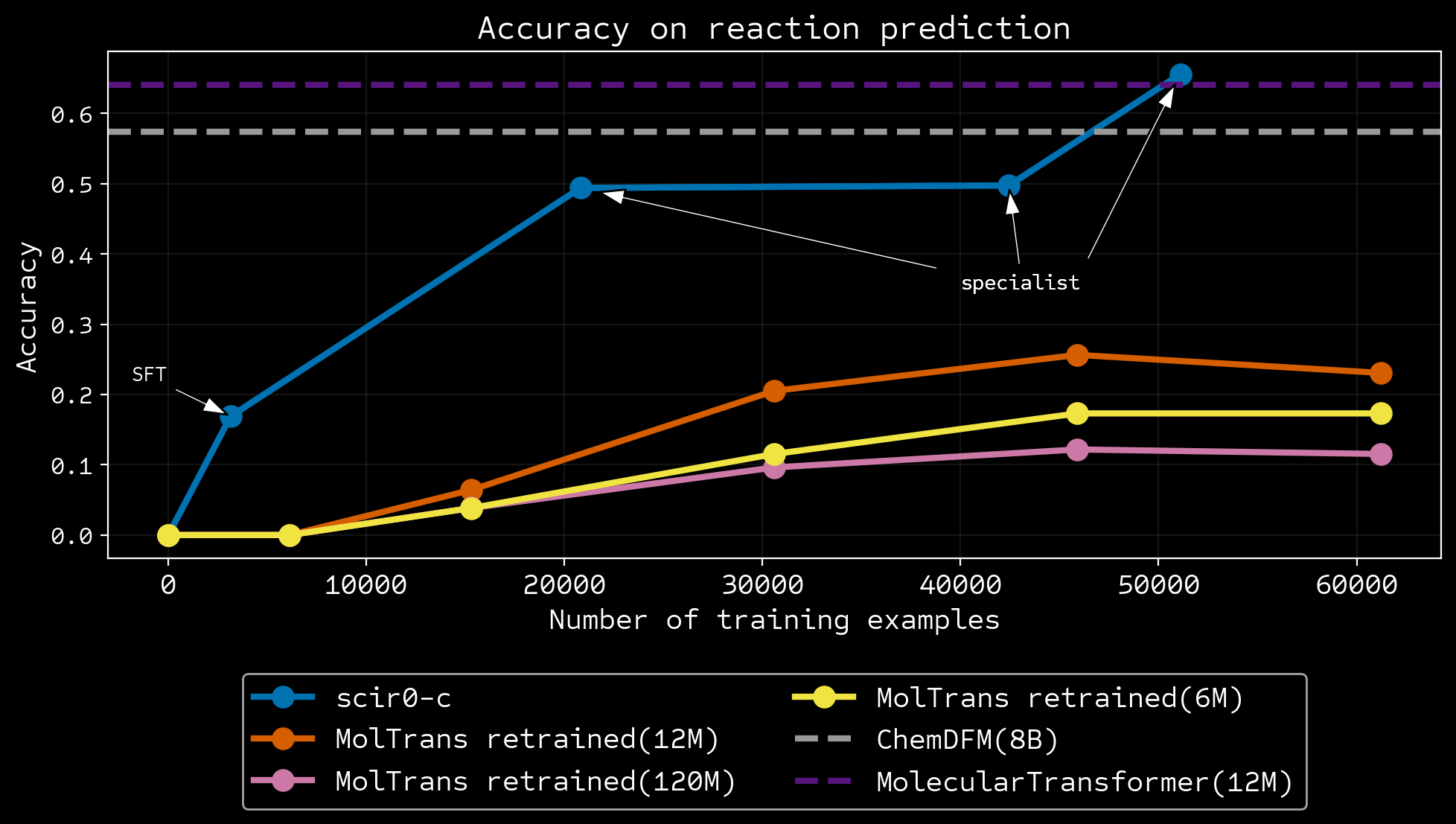
| reaction prediction | 1 | 61205 | molecule comparison | 10 | ORD |
| retrosynthesis | 1 | 67252 | ML model, database | 8 | mcule |
| BBB permeability | 2 | 2064 | multiple choice | 8 | BBB |
| pKa | 4 | 336 | multiple choice | 8 | IUPAC |
| safety | 11 | 5687 | multiple choice | 8 | Pubchem |
| molecular formula | 1 | 18738 | code | 10 | COCONUT |
| ADME | 12 | 1030 | multiple choice | 8 | Fang ADME |
| LD50 | 2 | 342 | multiple choice | 8 | Pubchem |
| Human receptor binding | 150 | 1663 | multiple choice | 8 | EveBio |
| property-regression-solubility | 2 | 464 | multiple choice | 8 | AqSolDB |
| property-regression-photo | 1 | 23 | multiple choice | 8 | Photoswitches |
| Total | 374 | 577790 | 8 | 81* | 12 |
Training Stages
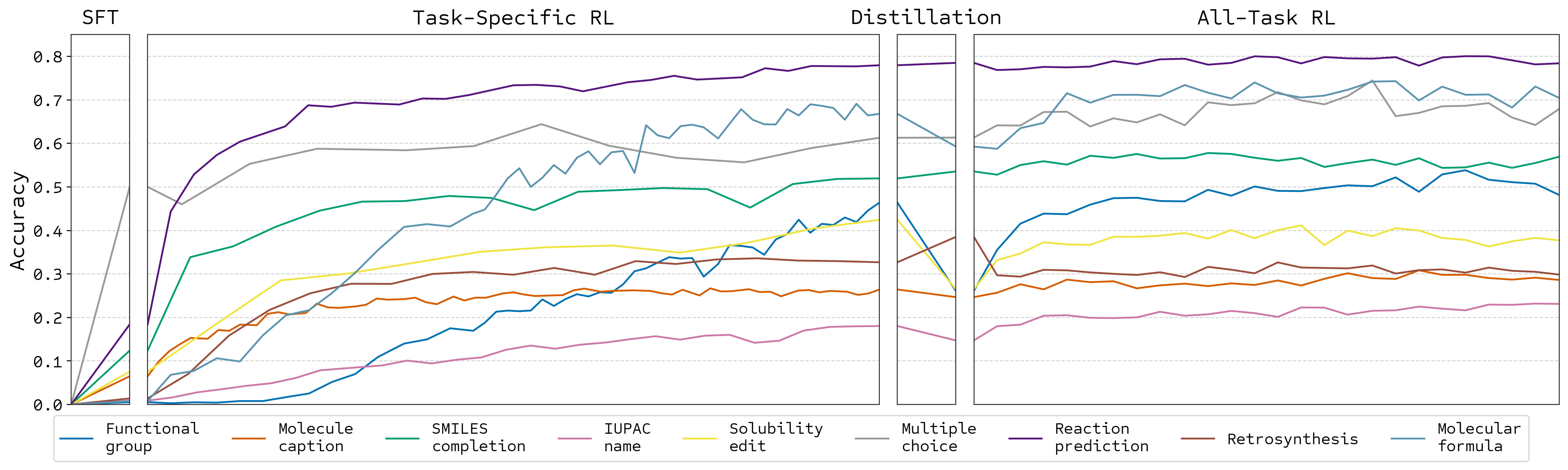
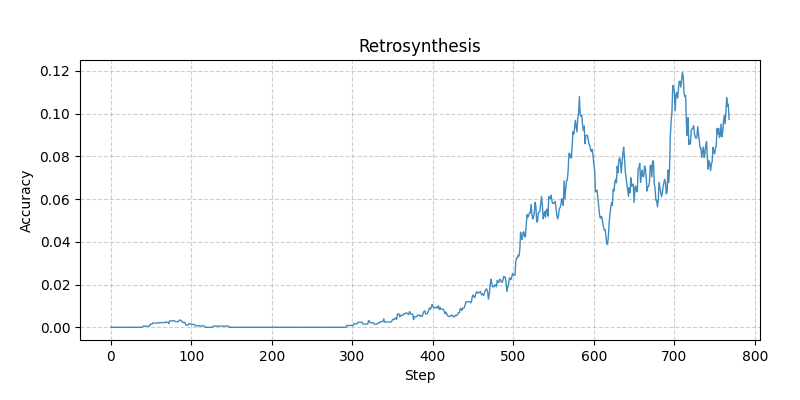
Can learn from zero accuracy
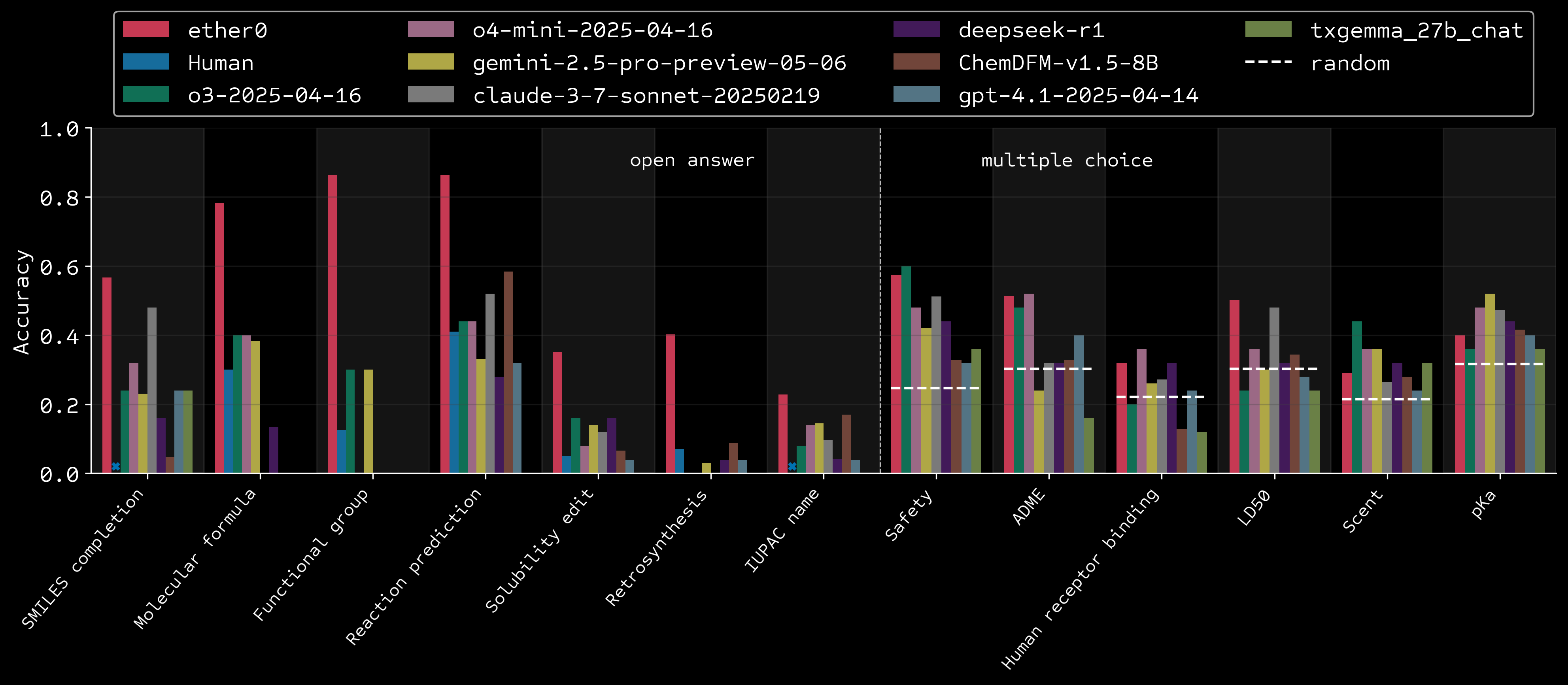
Results vs humans and frontier models
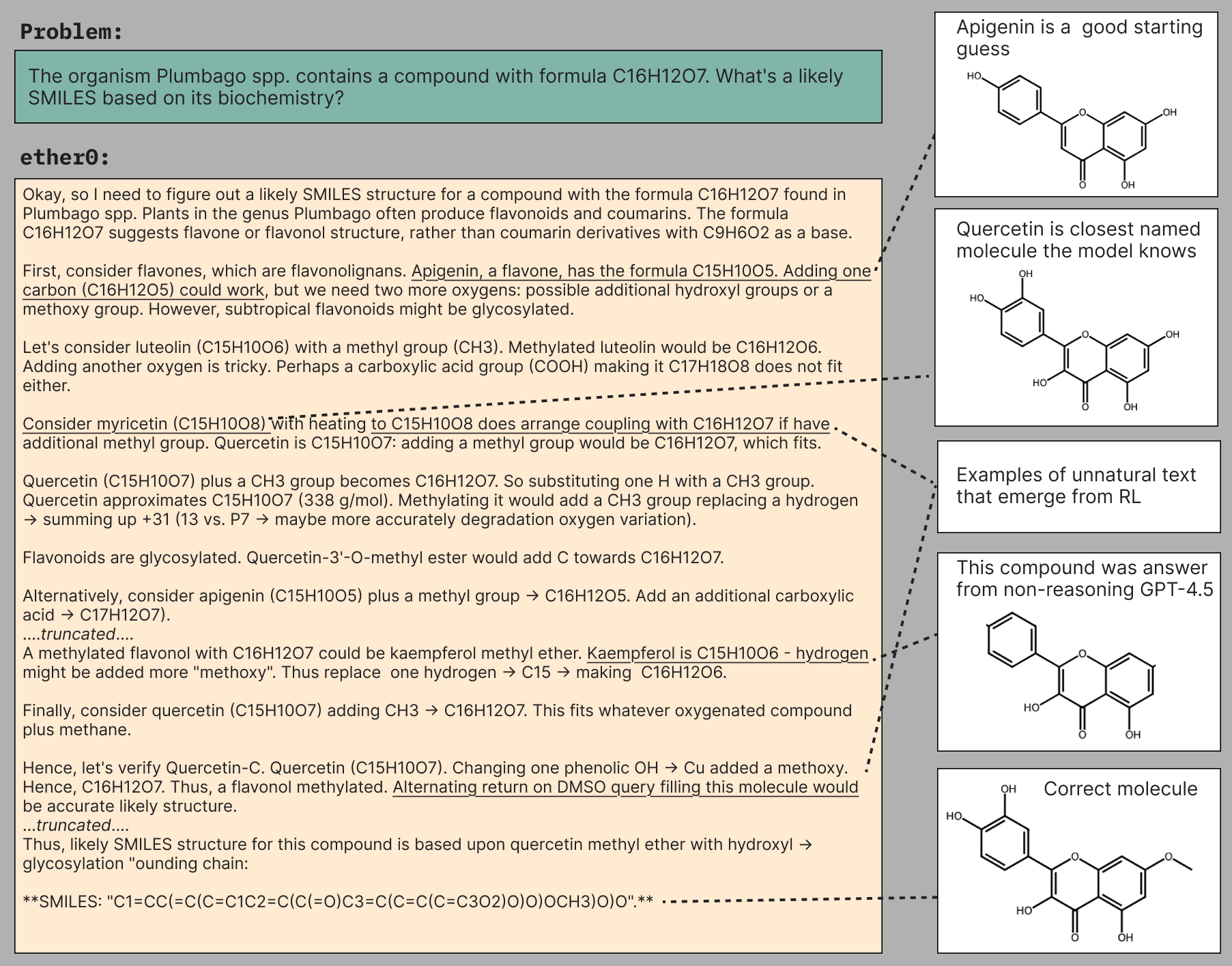
More data efficient
Acknowledgements for this talk
Rochester
- Sam Cox
FutureHouse
- Geemi Wellawatte
- Sam Rodriques
Edison Scientific
- Sid Narayanan
- James Braza
- Albert Bou
- Mayk Caldas
- Ludovico Michtner
- Mike Skarlinski
- Michael Pieler